Lineage for d3oeew2 (3oee W:83-357)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.37: P-loop containing nucleoside triphosphate hydrolases [52539] (1 superfamily)
3 layers: a/b/a, parallel or mixed beta-sheets of variable sizes
Superfamily c.37.1: P-loop containing nucleoside triphosphate hydrolases [52540] (27 families) 
division into families based on beta-sheet topologies
Family c.37.1.11: RecA protein-like (ATPase-domain) [52670] (22 proteins)
core: mixed beta-sheet of 8 strands, order 32451678; strand 7 is antiparallel to the rest
Protein Central domain of beta subunit of F1 ATP synthase [88779] (5 species)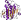 
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [310897] (6 PDB entries) 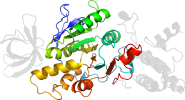
Domain d3oeew2: 3oee W:83-357 [239558]
Other proteins in same PDB: d3oeed1, d3oeed3, d3oeee1, d3oeee3, d3oeef1, d3oeef3, d3oeeg_, d3oeeh1, d3oeeh2, d3oeei_, d3oeem1, d3oeem3, d3oeen1, d3oeen3, d3oeeo1, d3oeeo3, d3oeep_, d3oeer_, d3oeev1, d3oeev3, d3oeew1, d3oeew3, d3oeex1, d3oeex3, d3oeey_
automated match to d2jdid3
complexed with anp, mg, po4; mutant
Details for d3oeew2
PDB Entry: 3oee (more details), 2.74 Å
PDB Description: Structure of four mutant forms of yeast F1 ATPase: alpha-F405S
PDB Compounds: (W:) ATP synthase subunit betaSCOPe Domain Sequences for d3oeew2:
Sequence; same for both SEQRES and ATOM records: (download)
>d3oeew2 c.37.1.11 (W:83-357) Central domain of beta subunit of F1 ATP synthase {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
isvpvgretlgriinvigepidergpiksklrkpihadppsfaeqstsaeiletgikvvd
llapyarggkiglfggagvgktvfiqelinniakahggfsvftgvgertregndlyremk
etgvinlegeskvalvfgqmneppgararvaltgltiaeyfrdeegqdvllfidnifrft
qagsevsallgripsavgyqptlatdmgllqeritttkkgsvtsvqavyvpaddltdpap
attfahldattvlsrgiselgiypavdpldsksrl
SCOPe Domain Coordinates for d3oeew2:
Click to download the PDB-style file with coordinates for d3oeew2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3oeew2:
- d3oeew2 first appeared in SCOPe 2.04
- d3oeew2 appears in SCOPe 2.07
 View in 3D View in 3DDomains from other chains: (mouse over for more information) d3oeed1, d3oeed2, d3oeed3, d3oeee1, d3oeee2, d3oeee3, d3oeef1, d3oeef2, d3oeef3, d3oeeg_, d3oeeh1, d3oeeh2, d3oeei_, d3oeem1, d3oeem2, d3oeem3, d3oeen1, d3oeen2, d3oeen3, d3oeeo1, d3oeeo2, d3oeeo3, d3oeep_, d3oeer_, d3oeev1, d3oeev2, d3oeev3, d3oeex1, d3oeex2, d3oeex3, d3oeey_ |