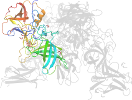
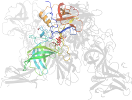
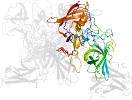Lineage for d6h6md_ (6h6m D:)
- Root: SCOPe 2.08

Class b: All beta proteins [48724] (180 folds) 
Fold b.121: Nucleoplasmin-like/VP (viral coat and capsid proteins) [88632] (7 superfamilies)
sandwich; 8 strands in 2 sheets; jelly-roll; some members can have additional 1-2 strands
characteristic interaction between the domains of this fold allows the formation of five-fold and pseudo six-fold assemblies
Superfamily b.121.4: Positive stranded ssRNA viruses [88633] (10 families) 
Family b.121.4.0: automated matches [191569] (1 protein)
not a true family
Protein automated matches [190988] (51 species)
not a true protein
Species Murine norovirus gv/cr10/2005/usa [TaxId:463724] [347267] (2 PDB entries) 
Domain d6h6md_: 6h6m D: [411202]
Other proteins in same PDB: d6h6me_, d6h6mf_
automated match to d5or7a_
complexed with cho, edo, na, peg, pge
Details for d6h6md_
PDB Entry: 6h6m (more details), 2.38 Å
PDB Description: cr10 murine norovirus protruding domain in complex with the cd300lf receptor and glycochenodeoxycholate (gcdca)
PDB Compounds: (D:) capsid proteinSCOPe Domain Sequences for d6h6md_:
Sequence; same for both SEQRES and ATOM records: (download)
>d6h6md_ b.121.4.0 (D:) automated matches {Murine norovirus gv/cr10/2005/usa [TaxId: 463724]}
rmvdlpvlqprlctharwpapiygllvdpslpsnpqwqngrvhvdgtllgttpvsgswvs
cfaaeaayefqsgtgevatftlieqdgsayvpgdraaplgypdfsgqleievqtettktg
dklkvttfemilgpttnvdqapyqgrvyasltavasldlvdgrvravprsiygfqdvipe
yndgllvplappigpflpgevllrfrtymrqldtadaaaeaidcalpqefiswfasnaft
vqsdalllryrntltgqllfecklysegyialsysgsgpltfptdgffevvswvprlfql
asv
SCOPe Domain Coordinates for d6h6md_:
Click to download the PDB-style file with coordinates for d6h6md_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d6h6md_:
- d6h6md_ is new in SCOPe 2.08-stable
 View in 3D
View in 3D