Lineage for d3oehi_ (3oeh I:)
- Root: SCOPe 2.08

Class a: All alpha proteins [46456] (290 folds) 
Fold a.137: Non-globular all-alpha subunits of globular proteins [48661] (14 superfamilies)
not a true fold
annotated by the SCOP(e) curators as 'not a true fold'
Superfamily a.137.8: Epsilon subunit of mitochondrial F1F0-ATP synthase [48690] (1 family) 
automatically mapped to Pfam PF04627
Family a.137.8.1: Epsilon subunit of mitochondrial F1F0-ATP synthase [48691] (2 proteins) 
Protein Epsilon subunit of mitochondrial F1F0-ATP synthase [48692] (2 species) 
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [310956] (5 PDB entries) 
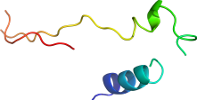
Domain d3oehi_: 3oeh I: [294218]
Other proteins in same PDB: d3oehd1, d3oehd2, d3oehd3, d3oehe1, d3oehe2, d3oehe3, d3oehf1, d3oehf2, d3oehf3, d3oehg_, d3oehh1, d3oehh2, d3oehm1, d3oehm2, d3oehm3, d3oehn1, d3oehn2, d3oehn3, d3oeho1, d3oeho2, d3oeho3, d3oehp_, d3oehv1, d3oehv2, d3oehv3, d3oehw1, d3oehw2, d3oehw3, d3oehx1, d3oehx2, d3oehx3
automated match to d2hldi_
complexed with anp, mg; mutant
Details for d3oehi_
PDB Entry: 3oeh (more details), 3 Å
PDB Description: Structure of four mutant forms of yeast F1 ATPase: beta-V279F
PDB Compounds: (I:) ATP synthase subunit epsilonSCOPe Domain Sequences for d3oehi_:
Sequence, based on SEQRES records: (download)
>d3oehi_ a.137.8.1 (I:) Epsilon subunit of mitochondrial F1F0-ATP synthase {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
isyaaylnvaaqairsslktelqtasvlnrsqtdafytqykngtaaseptpitk
Sequence, based on observed residues (ATOM records): (download)
>d3oehi_ a.137.8.1 (I:) Epsilon subunit of mitochondrial F1F0-ATP synthase {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
isyaaylnvaaqairsktelqtasvlnrsqtdafytqyknaseptpitk
SCOPe Domain Coordinates for d3oehi_:
Click to download the PDB-style file with coordinates for d3oehi_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3oehi_:
- d3oehi_ first appeared in SCOPe 2.06
- d3oehi_ appears in SCOPe 2.07
 View in 3D View in 3DDomains from other chains: (mouse over for more information) d3oehd1, d3oehd2, d3oehd3, d3oehe1, d3oehe2, d3oehe3, d3oehf1, d3oehf2, d3oehf3, d3oehg_, d3oehh1, d3oehh2, d3oehm1, d3oehm2, d3oehm3, d3oehn1, d3oehn2, d3oehn3, d3oeho1, d3oeho2, d3oeho3, d3oehp_, d3oehr_, d3oehv1, d3oehv2, d3oehv3, d3oehw1, d3oehw2, d3oehw3, d3oehx1, d3oehx2, d3oehx3 |