Lineage for d4e6pd_ (4e6p D:)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.2: NAD(P)-binding Rossmann-fold domains [51734] (1 superfamily)
core: 3 layers, a/b/a; parallel beta-sheet of 6 strands, order 321456
The nucleotide-binding modes of this and the next two folds/superfamilies are similar
Superfamily c.2.1: NAD(P)-binding Rossmann-fold domains [51735] (13 families) 
Family c.2.1.2: Tyrosine-dependent oxidoreductases [51751] (73 proteins)
also known as short-chain dehydrogenases and SDR family
parallel beta-sheet is extended by 7th strand, order 3214567; left-handed crossover connection between strands 6 and 7
has additional subdomain(s) that are not in the common domain

Protein automated matches [190085] (59 species)
not a true protein
Species Sinorhizobium meliloti [TaxId:266834] [195628] (1 PDB entry) 
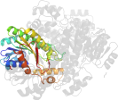
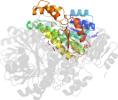
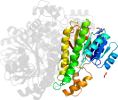
Domain d4e6pd_: 4e6p D: [195630]
automated match to d1k2wa_
complexed with edo, na
Details for d4e6pd_
PDB Entry: 4e6p (more details), 2.1 Å
PDB Description: Crystal structure of a probable sorbitol dehydrogenase (Target PSI-012078) from Sinorhizobium meliloti 1021
PDB Compounds: (D:) Probable sorbitol dehydrogenase (L-iditol 2-dehydrogenase)SCOPe Domain Sequences for d4e6pd_:
Sequence; same for both SEQRES and ATOM records: (download)
>d4e6pd_ c.2.1.2 (D:) automated matches {Sinorhizobium meliloti [TaxId: 266834]}
krlegksalitgsargigrafaeayvregatvaiadidierarqaaaeigpaayavqmdv
trqdsidaaiaatvehaggldilvnnaalfdlapiveitresyeklfainvagtlftlqa
aarqmiaqgrggkiinmasqagrrgealvaiycatkaavisltqsagldlikhrinvnai
apgvvdgehwdgvdalfaryenrprgekkrlvgeavpfgrmgtaedltgmaiflasaesd
yivsqtynvdggnwms
SCOPe Domain Coordinates for d4e6pd_:
Click to download the PDB-style file with coordinates for d4e6pd_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d4e6pd_:
- d4e6pd_ first appeared in SCOPe 2.02
- d4e6pd_ appears in SCOPe 2.07

 View in 3D
View in 3D