Lineage for d1iqua1 (1iqu A:172-416)
- Root: SCOP 1.61

Class a: All alpha proteins [46456] (151 folds) 
Fold a.99: FAD-binding (C-terminal) domain of DNA photolyase [48172] (1 superfamily) 
Superfamily a.99.1: FAD-binding (C-terminal) domain of DNA photolyase [48173] (1 family) 

Family a.99.1.1: FAD-binding (C-terminal) domain of DNA photolyase [48174] (1 protein) 
Protein FAD-binding (C-terminal) domain of DNA photolyase [48175] (3 species) 
Species Thermus thermophilus [TaxId:274] [69083] (2 PDB entries) 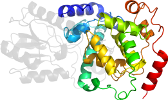
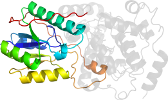
Domain d1iqua1: 1iqu A:172-416 [71280]
Other proteins in same PDB: d1iqua2
Details for d1iqua1
PDB Entry: 1iqu (more details), 2.2 Å
PDB Description: Crystal structure of photolyase-thymine complex
SCOP Domain Sequences for d1iqua1:
Sequence; same for both SEQRES and ATOM records: (download)
>d1iqua1 a.99.1.1 (A:172-416) FAD-binding (C-terminal) domain of DNA photolyase {Thermus thermophilus}
lplpepgeeaalaglrafleaklpryaeerdrldgeggsrlspyfalgvlsprlaaweae
rrggegarkwvaellwrdfsyhllyhfpwmaerpldprfqafpwqedealfqawyegktg
vplvdaamrelhatgflsnrarmnaaqfavkhlllpwkrceeafrhllldgdravnlqgw
qwagglgvdaapyfrvfnpvlqgerhdpegrwlkrwapeypsyapkdpvvdleearrryl
rlard
SCOP Domain Coordinates for d1iqua1:
Click to download the PDB-style file with coordinates for d1iqua1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1iqua1:
- d1iqua1 is new in SCOP 1.61
- d1iqua1 appears in SCOP 1.63
- d1iqua1 appears in the current release, SCOPe 2.08
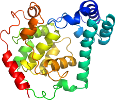

 View in 3D
View in 3D