Lineage for d1clib1 (1cli B:1021-1170)
- Root: SCOPe 2.01

Class d: Alpha and beta proteins (a+b) [53931] (376 folds) 
Fold d.79: Bacillus chorismate mutase-like [55297] (9 superfamilies)
core: beta-alpha-beta-alpha-beta(2); mixed beta-sheet: order: 1423, strand 4 is antiparallel to the rest
Superfamily d.79.4: PurM N-terminal domain-like [55326] (1 family) 

Family d.79.4.1: PurM N-terminal domain-like [55327] (6 proteins)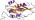 
Protein Aminoimidazole ribonucleotide synthetase (PurM) N-terminal domain [55328] (1 species) 
Species Escherichia coli [TaxId:562] [55329] (1 PDB entry) 
Domain d1clib1: 1cli B:1021-1170 [39817]
Other proteins in same PDB: d1clia2, d1clib2, d1clic2, d1clid2
complexed with so4
Details for d1clib1
PDB Entry: 1cli (more details), 2.5 Å
PDB Description: x-ray crystal structure of aminoimidazole ribonucleotide synthetase (purm), from the e. coli purine biosynthetic pathway, at 2.5 a resolution
PDB Compounds: (B:) protein (phosphoribosyl-aminoimidazole synthetase)SCOPe Domain Sequences for d1clib1:
Sequence; same for both SEQRES and ATOM records: (download)
>d1clib1 d.79.4.1 (B:1021-1170) Aminoimidazole ribonucleotide synthetase (PurM) N-terminal domain {Escherichia coli [TaxId: 562]}
alvgrikgvvkktrrpevmgglggfgalcalpqkyrepvlvsgtdgvgtklrlamdlkrh
dtigidlvamcvndlvvqgaeplffldyyatgkldvdtasavisgiaegclqsgcslvgg
etaempgmyhgedydvagfcvgvvekseii
SCOPe Domain Coordinates for d1clib1:
Click to download the PDB-style file with coordinates for d1clib1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1clib1:
- d1clib1 first appeared (with stable ids) in SCOP 1.55
- d1clib1 appears in SCOP 1.75
- d1clib1 appears in SCOPe 2.02
- d1clib1 appears in the current release, SCOPe 2.08
 View in 3D
View in 3D