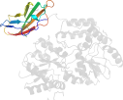Lineage for d3csba2 (3csb A:5-370)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.94: Periplasmic binding protein-like II [53849] (1 superfamily)
consists of two similar intertwined domain with 3 layers (a/b/a) each: duplication
mixed beta-sheet of 5 strands, order 21354; strand 5 is antiparallel to the rest
Superfamily c.94.1: Periplasmic binding protein-like II [53850] (4 families) 
Similar in architecture to the superfamily I but partly differs in topology
Family c.94.1.1: Phosphate binding protein-like [53851] (45 proteins)
has additional insertions and/or extensions that are not grouped together
Protein D-maltodextrin-binding protein, MBP [53862] (5 species)
contains a few additional helices in the C-terminal extension; homologous to thiaminase I
Species Escherichia coli K-12 [TaxId:83333] [159806] (14 PDB entries) 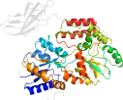
Domain d3csba2: 3csb A:5-370 [156960]
Other proteins in same PDB: d3csba1
automatically matched to d1mpba_
complexed with 1pe, edo, mn, peg, pg4, pge
has additional insertions and/or extensions that are not grouped together
Details for d3csba2
PDB Entry: 3csb (more details), 2 Å
PDB Description: crystal structure of monobody ysx1/maltose binding protein fusion complex
PDB Compounds: (A:) Maltose-binding protein Monobody YSX1 FusionSCOPe Domain Sequences for d3csba2:
Sequence; same for both SEQRES and ATOM records: (download)
>d3csba2 c.94.1.1 (A:5-370) D-maltodextrin-binding protein, MBP {Escherichia coli K-12 [TaxId: 83333]}
gklviwingdkgynglaevgkkfekdtgikvtvehpdkleekfpqvaatgdgpdiifwah
drfggyaqsgllaeitpdkafqdklypftwdavryngkliaypiavealsliynkdllpn
ppktweeipaldkelkakgksalmfnlqepyftwpliaadggyafkyengkydikdvgvd
nagakagltflvdliknkhmnadtdysiaeaafnkgetamtingpwawsnidtskvnygv
tvlptfkgqpskpfvgvlsaginaaspnkelakeflenylltdegleavnkdkplgaval
ksyeeelakdpriaatmenaqkgeimpnipqmsafwyavrtavinaasgrqtvdealkda
qtritk
SCOPe Domain Coordinates for d3csba2:
Click to download the PDB-style file with coordinates for d3csba2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3csba2:
- d3csba2 first appeared in SCOP 1.75
- d3csba2 appears in SCOPe 2.07

 View in 3D
View in 3D