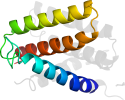
Lineage for d2p06b1 (2p06 B:1-83)
- Root: SCOPe 2.08

Class a: All alpha proteins [46456] (290 folds) 
Fold a.204: all-alpha NTP pyrophosphatases [101385] (1 superfamily)
multihelical: dimeric 4-helical bundle surrounded by other helices; oligomerizes further in a tetramer
Superfamily a.204.1: all-alpha NTP pyrophosphatases [101386] (5 families) 
basic module consist of 5 active site-forming helices; four from one subunit/structural repeat; the fifth from the other subunit/repeat
Family a.204.1.3: AF0060-like [140794] (1 protein)
consists of a single member with an orphan sequence; "circular permutation" of putative active site residues: the catalytic lysine resides in the N-terminal helix instead of the missing in this family C-terminal helix; probable biological unit is a tertramer, distinct from the MazG family tetramer
Protein Hypothetical protein AF0060 [140795] (1 species)
Uniprot O30176 1-84
Species Archaeoglobus fulgidus [TaxId:2234] [140796] (1 PDB entry) 
Domain d2p06b1: 2p06 B:1-83 [139453]
Other proteins in same PDB: d2p06b2
complexed with gol, mg
Details for d2p06b1
PDB Entry: 2p06 (more details), 2.1 Å
PDB Description: Crystal structure of a predicted coding region AF_0060 from Archaeoglobus fulgidus DSM 4304
PDB Compounds: (B:) Hypothetical protein AF_0060SCOPe Domain Sequences for d2p06b1:
Sequence; same for both SEQRES and ATOM records: (download)
>d2p06b1 a.204.1.3 (B:1-83) Hypothetical protein AF0060 {Archaeoglobus fulgidus [TaxId: 2234]}
mdyfrlaekflremhakymkrvsrpgntprpwfdfseerllsrlfeemdelreavekedw
enlrdelldvanfcmylwgklsv
SCOPe Domain Coordinates for d2p06b1:
Click to download the PDB-style file with coordinates for d2p06b1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d2p06b1:
- d2p06b1 first appeared in SCOP 1.73
- d2p06b1 appears in SCOPe 2.07
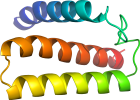
 View in 3D
View in 3D