Lineage for d2akmb1 (2akm B:140-432)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.1: TIM beta/alpha-barrel [51350] (34 superfamilies)
contains parallel beta-sheet barrel, closed; n=8, S=8; strand order 12345678
the first seven superfamilies have similar phosphate-binding sites
Superfamily c.1.11: Enolase C-terminal domain-like [51604] (3 families) 
binds metal ion (magnesium or manganese) in conserved site inside barrel
N-terminal alpha+beta domain is common to this superfamily
Family c.1.11.1: Enolase [51605] (2 proteins)
automatically mapped to Pfam PF00113
Protein Enolase [51606] (11 species)
Fold of this protein slightly differs from common fold in topology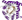
Species Human (Homo sapiens), gamma isoform [TaxId:9606] [110371] (15 PDB entries)
Uniprot P09104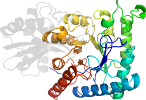
Domain d2akmb1: 2akm B:140-432 [126921]
Other proteins in same PDB: d2akma2, d2akmb2
automated match to d1te6a1
complexed with mg, po4, trs
Details for d2akmb1
PDB Entry: 2akm (more details), 1.92 Å
PDB Description: Fluoride Inhibition of Enolase: Crystal Structure of the Inhibitory Complex
PDB Compounds: (B:) Gamma enolaseSCOPe Domain Sequences for d2akmb1:
Sequence; same for both SEQRES and ATOM records: (download)
>d2akmb1 c.1.11.1 (B:140-432) Enolase {Human (Homo sapiens), gamma isoform [TaxId: 9606]}
sdlilpvpafnvinggshagnklamqefmilpvgaesfrdamrlgaevyhtlkgvikdky
gkdatnvgdeggfapnilensealelvkeaidkagytekivigmdvaasefyrdgkydld
fksptdpsryitgdqlgalyqdfvrdypvvsiedpfdqddwaawskftanvgiqivgddl
tvtnpkrieraveekacnclllkvnqigsvteaiqacklaqengwgvmvshrsgetedtf
iadlvvglctgqiktgapcrserlakynqlmrieeelgdearfaghnfrnpsv
SCOPe Domain Coordinates for d2akmb1:
Click to download the PDB-style file with coordinates for d2akmb1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d2akmb1:
- d2akmb1 first appeared in SCOP 1.73
- d2akmb1 appears in SCOPe 2.07
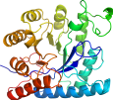
 View in 3D
View in 3D