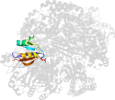
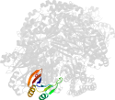
Lineage for d1is7g_ (1is7 G:)
- Root: SCOP 1.67

Class d: Alpha and beta proteins (a+b) [53931] (260 folds) 
Fold d.96: T-fold [55619] (1 superfamily)
beta(2)-alpha(2)-beta(2); 2 layers: alpha/beta; antiparallel sheet 1234
tunnel-shaped: its known members form wide oligomeric barrels different sizes
Superfamily d.96.1: Tetrahydrobiopterin biosynthesis enzymes-like [55620] (4 families) 
bind purine or pterin in topologically similar sites between subunits
Family d.96.1.1: GTP cyclohydrolase I [55621] (1 protein) 
Protein GTP cyclohydrolase I [55622] (3 species)
beta-sheets of five subunits form a barrel, closed: n=20, S=20
Species Rat (Rattus norvegicus) [TaxId:10116] [69790] (2 PDB entries) 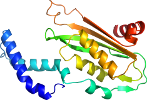
Domain d1is7g_: 1is7 G: [66307]
Other proteins in same PDB: d1is7k_, d1is7l_, d1is7m_, d1is7n_, d1is7o_, d1is7p_, d1is7q_, d1is7r_, d1is7s_, d1is7t_
Details for d1is7g_
PDB Entry: 1is7 (more details), 2.8 Å
PDB Description: crystal structure of rat gtpchi/gfrp stimulatory complex
SCOP Domain Sequences for d1is7g_:
Sequence; same for both SEQRES and ATOM records: (download)
>d1is7g_ d.96.1.1 (G:) GTP cyclohydrolase I {Rat (Rattus norvegicus)}
rprseednelnlpnlaaayssilrslgedpqrqgllktpwraatamqfftkgyqetisdv
lndaifdedhdemvivkdidmfsmcehhlvpfvgrvhigylpnkqvlglsklariveiys
rrlqvqerltkqiavaitealqpagvgvvieathmcmvmrgvqkmnsktvtstmlgvfre
dpktreefltlirs
SCOP Domain Coordinates for d1is7g_:
Click to download the PDB-style file with coordinates for d1is7g_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1is7g_:
- d1is7g_ first appeared in SCOP 1.59
- d1is7g_ appears in SCOP 1.65
- d1is7g_ appears in SCOP 1.69
- d1is7g_ appears in the current release, SCOPe 2.08
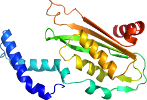
 View in 3D
View in 3D