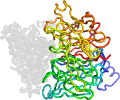
Lineage for d2madh_ (2mad H:)
- Root: SCOPe 2.08

Class b: All beta proteins [48724] (180 folds) 
Fold b.69: 7-bladed beta-propeller [50964] (15 superfamilies)
consists of seven 4-stranded beta-sheet motifs; meander
Superfamily b.69.2: YVTN repeat-like/Quinoprotein amine dehydrogenase [50969] (4 families) 
Family b.69.2.1: Methylamine dehydrogenase, H-chain [50970] (1 protein)
less regular propeller without notable sequence repeats; quinone cofactor is a tryptophan derivative in another subunit
automatically mapped to Pfam PF06433
this is a repeat family; one repeat unit is 1mg2 A:132-178 found in domain
Protein Methylamine dehydrogenase, H-chain [50971] (2 species) 
Species Paracoccus versutus (Thiobacillus versutus) [TaxId:34007] [50973] (3 PDB entries) 
Domain d2madh_: 2mad H: [27639]
Other proteins in same PDB: d2madl_
Details for d2madh_
PDB Entry: 2mad (more details), 2.25 Å
PDB Description: the active site structure of methylamine dehydrogenase: hydrazines identify c6 as the reactive site of the tryptophan derived quinone cofactor
PDB Compounds: (H:) methylamine dehydrogenase (heavy subunit)SCOPe Domain Sequences for d2madh_:
Sequence; same for both SEQRES and ATOM records: (download)
>d2madh_ b.69.2.1 (H:) Methylamine dehydrogenase, H-chain {Paracoccus versutus (Thiobacillus versutus) [TaxId: 34007]}
ssasaaaaaaaaalaagaadgptndeapgadgrrsyinlpahhsaiiqqwvldagsgsil
ghvnggflpnpvaahsgsefalastsfsriakgkrtdyvevfdpvtflpiadielpdapr
fdvgpyswmnantpnnadllffqfaagpavglvvqggssddqllssptcyhihpgapstf
yllcaqgglaktdhaggaagaglvgamltaaqnlltqpaqanksgrivwpvysgkilqad
isaagatnkapidalsggrkadtwrpggwqqvaylkssdgiylltseqsawklhaaakev
tsvtglvgqtssqislghdvdaisvaqdggpdlyalsagtevlhiydagagdqdqstvel
gsgpqvlsvmnea
SCOPe Domain Coordinates for d2madh_:
Click to download the PDB-style file with coordinates for d2madh_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d2madh_:
- d2madh_ first appeared (with stable ids) in SCOP 1.55
- d2madh_ appears in SCOPe 2.07
 View in 3D
View in 3D