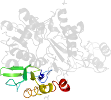Lineage for d3mjfa2 (3mjf A:104-327)
- Root: SCOPe 2.04

Class d: Alpha and beta proteins (a+b) [53931] (380 folds) 
Fold d.142: ATP-grasp [56058] (2 superfamilies)
Consists of two subdomains with different alpha+beta folds
shares functional and structural similarities with the PIPK and protein kinase superfamilies
Superfamily d.142.1: Glutathione synthetase ATP-binding domain-like [56059] (10 families) 
Family d.142.1.2: BC ATP-binding domain-like [56067] (7 proteins) 
Protein automated matches [254674] (2 species)
not a true protein
Species Yersinia pestis [TaxId:214092] [255970] (1 PDB entry) 
Domain d3mjfa2: 3mjf A:104-327 [247725]
Other proteins in same PDB: d3mjfa1, d3mjfa3
automated match to d1gsoa3
complexed with bme, edo, gol, na, peg, pge, so4
Details for d3mjfa2
PDB Entry: 3mjf (more details), 1.47 Å
PDB Description: phosphoribosylamine-glycine ligase from yersinia pestis
PDB Compounds: (A:) Phosphoribosylamine--glycine ligaseSCOPe Domain Sequences for d3mjfa2:
Sequence; same for both SEQRES and ATOM records: (download)
>d3mjfa2 d.142.1.2 (A:104-327) automated matches {Yersinia pestis [TaxId: 214092]}
skaftkdflarhnipsaeyqnftdveaalayvrqkgapivikadglaagkgvivamtqee
aetavndmlagnafgdaghrivveefldgeeasfivmvdgenvlpmatsqdhkrvgdgdt
gpntggmgayspapvvtddvhqrvmdqviwptvrgmaaegniytgflyaglmisadgqpk
viefncrfgdpetqpimlrmrsdlvelclagtqgklnektsdwd
SCOPe Domain Coordinates for d3mjfa2:
Click to download the PDB-style file with coordinates for d3mjfa2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3mjfa2:
- d3mjfa2 is new in SCOPe 2.04-stable
- d3mjfa2 appears in SCOPe 2.05
- d3mjfa2 appears in the current release, SCOPe 2.08
 View in 3D
View in 3D