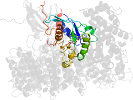
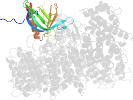Lineage for d4havb_ (4hav B:)
- Root: SCOPe 2.06

Class b: All beta proteins [48724] (177 folds) 
Fold b.55: PH domain-like barrel [50728] (2 superfamilies)
barrel, partly opened; n*=6, S*=12; meander; capped by an alpha-helix
Superfamily b.55.1: PH domain-like [50729] (14 families) 
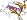

Family b.55.1.0: automated matches [191311] (1 protein)
not a true family
Protein automated matches [190052] (6 species)
not a true protein
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [194506] (31 PDB entries) 
Domain d4havb_: 4hav B: [222462]
Other proteins in same PDB: d4hava_
automated match to d4hb3b_
complexed with aa8, cl, gnp, mg
Details for d4havb_
PDB Entry: 4hav (more details), 2 Å
PDB Description: crystal structure of crm1 inhibitor anguinomycin a in complex with crm1-ran-ranbp1
PDB Compounds: (B:) Ran-specific GTPase-activating protein 1SCOPe Domain Sequences for d4havb_:
Sequence, based on SEQRES records: (download)
>d4havb_ b.55.1.0 (B:) automated matches {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
ihfepvvhlekvdvktmeedeevlykvraklfrfdkdakewkergtgdckflknkktnkv
rilmrrdktlkicanhiiapeytlkpnvgsdrswvyactadiaegeaeaftfairfgske
nadkfkeefekaqeinkk
Sequence, based on observed residues (ATOM records): (download)
>d4havb_ b.55.1.0 (B:) automated matches {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
ihfepvvktmeedeevlykvraklfrfdkdakewkergtgdckflknkktnkvrilmrrd
ktlkicanhiiapeytlkpnvgsdrswvyactadiaegeaeaftfairfgskenadkfke
efekaqeinkk
SCOPe Domain Coordinates for d4havb_:
Click to download the PDB-style file with coordinates for d4havb_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d4havb_:
- d4havb_ first appeared in SCOPe 2.03
- d4havb_ appears in SCOPe 2.05
- d4havb_ appears in SCOPe 2.07
- d4havb_ appears in the current release, SCOPe 2.08

 View in 3D
View in 3D