Lineage for d1xygc2 (1xyg C:163-326)
- Root: SCOPe 2.08

Class d: Alpha and beta proteins (a+b) [53931] (396 folds) 
Fold d.81: FwdE/GAPDH domain-like [55346] (4 superfamilies)
core: alpha-beta-alpha-beta(3); mixed sheet: 2134, strand 2 is parallel to strand 1
Superfamily d.81.1: Glyceraldehyde-3-phosphate dehydrogenase-like, C-terminal domain [55347] (5 families) 
N-terminal domain is the classic Rossmann-fold
Family d.81.1.1: GAPDH-like [55348] (6 proteins)
has many additional secondary structures
Protein N-acetyl-gamma-glutamyl-phosphate reductase ArgC [111046] (3 species) 
Species Thale cress (Arabidopsis thaliana) [TaxId:3702] [118038] (2 PDB entries)
Uniprot O82187 15-359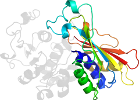
Domain d1xygc2: 1xyg C:163-326 [116226]
Other proteins in same PDB: d1xyga1, d1xygb1, d1xygc1, d1xygd1
Details for d1xygc2
PDB Entry: 1xyg (more details), 2.19 Å
PDB Description: x-ray structure of gene product from arabidopsis thaliana at2g19940
PDB Compounds: (C:) putative N-acetyl-gamma-glutamyl-phosphate reductaseSCOPe Domain Sequences for d1xygc2:
Sequence; same for both SEQRES and ATOM records: (download)
>d1xygc2 d.81.1.1 (C:163-326) N-acetyl-gamma-glutamyl-phosphate reductase ArgC {Thale cress (Arabidopsis thaliana) [TaxId: 3702]}
cypttiqlplvpllkanlikheniiidaksgvsgagrgakeanlyseiaegissygvtrh
rhvpeieqglsdvaqskvtvsftphlmpmirgmqstiyvemapgvrtedlhqqlktsyed
eefvkvldegvvprthnvrgsnychmsvfpdripgraiiisvid
SCOPe Domain Coordinates for d1xygc2:
Click to download the PDB-style file with coordinates for d1xygc2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1xygc2:
- d1xygc2 first appeared in SCOP 1.71
- d1xygc2 appears in SCOPe 2.07
 View in 3D
View in 3D