Lineage for d1upsa2 (1ups A:290-420)
- Root: SCOPe 2.08

Class b: All beta proteins [48724] (180 folds) 
Fold b.42: beta-Trefoil [50352] (8 superfamilies)
barrel, closed; n=6, S=12; and a hairpin triplet; meander
duplication: has internal pseudo threefold symmetry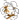
Superfamily b.42.2: Ricin B-like lectins [50370] (4 families) 

Family b.42.2.3: GlcNAc-alpha-1,4-Gal-releasing endo-beta-galactosidase, GngC, C-terminal domain [117212] (1 protein) 
Protein GlcNAc-alpha-1,4-Gal-releasing endo-beta-galactosidase, GngC, C-terminal domain [117213] (1 species) 
Species Clostridium perfringens [TaxId:1502] [117214] (1 PDB entry)
Uniprot Q934G8 17-420
Domain d1upsa2: 1ups A:290-420 [113395]
Other proteins in same PDB: d1upsa1, d1upsb1
complexed with ca
Details for d1upsa2
PDB Entry: 1ups (more details), 1.82 Å
PDB Description: glcnac[alpha]1-4gal releasing endo-[beta]-galactosidase from clostridium perfringens
PDB Compounds: (A:) glcnac-alpha-1,4-gal-releasing endo-beta-galactosidaseSCOPe Domain Sequences for d1upsa2:
Sequence; same for both SEQRES and ATOM records: (download)
>d1upsa2 b.42.2.3 (A:290-420) GlcNAc-alpha-1,4-Gal-releasing endo-beta-galactosidase, GngC, C-terminal domain {Clostridium perfringens [TaxId: 1502]}
nnylirnrqtgkflyieenndkvsygditlkneknakwskeyrdgytllknnetgeylni
enqtgyiehgkvpktwwsaqwsevpvdgytrfvnrwkpnmsihtesyegvlqygnvpnty
wtsqwqlipve
SCOPe Domain Coordinates for d1upsa2:
Click to download the PDB-style file with coordinates for d1upsa2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1upsa2:
- d1upsa2 first appeared in SCOP 1.71
- d1upsa2 appears in SCOPe 2.07

 View in 3D
View in 3D