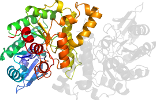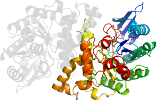Lineage for d6mgta_ (6mgt A:)
- Root: SCOPe 2.07

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.1: TIM beta/alpha-barrel [51350] (33 superfamilies)
contains parallel beta-sheet barrel, closed; n=8, S=8; strand order 12345678
the first seven superfamilies have similar phosphate-binding sites
Superfamily c.1.9: Metallo-dependent hydrolases [51556] (19 families) 
the beta-sheet barrel is similarly distorted and capped by a C-terminal helix
has transition metal ions bound inside the barrel
Family c.1.9.15: PP1699/LP2961-like [141819] (6 proteins)
Pfam PF04909; Amidohydrolase; stand-alone domain
Protein 2-amino-3-carboxymuconate 6-semialdehyde decarboxylase NbaD [141824] (1 species) 
Species Pseudomonas fluorescens [TaxId:294] [141825] (14 PDB entries)
Uniprot Q83V25 3-333
Domain d6mgta_: 6mgt A: [370601]
automated match to d4eraa_
complexed with co; mutant
Details for d6mgta_
PDB Entry: 6mgt (more details), 2.77 Å
PDB Description: crystal structure of alpha-amino-beta-carboxymuconate-epsilon- semialdehyde decarboxylase mutant h110a
PDB Compounds: (A:) 2-amino-3-carboxymuconate 6-semialdehyde decarboxylaseSCOPe Domain Sequences for d6mgta_:
Sequence; same for both SEQRES and ATOM records: (download)
>d6mgta_ c.1.9.15 (A:) 2-amino-3-carboxymuconate 6-semialdehyde decarboxylase NbaD {Pseudomonas fluorescens [TaxId: 294]}
pridmhshffpriseqeaakfdanhapwlqvsakgdtgsimmgknnfrpvyqalwdpafr
ieemdaqgvdvqvtcatpvmfgytweankaaqwaermndfalefaaanpqrikvlaqvpl
qdldlackeasravaaghlgiqignhlgdkdlddatleaflthcanedipilvhpwdmmg
gqrmkkwmlpwlvampaetqlailslilsgaferipkslkicfghgggsfafllgrvdna
wrhrdivredcprppseyvdrffvdsavfnpgalellvsvmgedrvmlgsdypfplgeqk
igglvlssnlgesakdkiisgnaskffnin
SCOPe Domain Coordinates for d6mgta_:
Click to download the PDB-style file with coordinates for d6mgta_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d6mgta_:
- d6mgta_ is new in SCOPe 2.07-stable
- d6mgta_ appears in the current release, SCOPe 2.08

 View in 3D
View in 3D