Lineage for d1nifa2 (1nif A:167-340)
- Root: SCOPe 2.07

Class b: All beta proteins [48724] (178 folds) 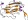
Fold b.6: Cupredoxin-like [49502] (2 superfamilies)
sandwich; 7 strands in 2 sheets, greek-key
variations: some members have additional 1-2 strands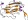
Superfamily b.6.1: Cupredoxins [49503] (8 families) 
contains copper-binding site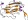
Family b.6.1.3: Multidomain cupredoxins [49550] (8 proteins) 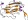
Protein Nitrite reductase, NIR [49551] (5 species)
consists of two domains of this fold
Species Achromobacter cycloclastes [TaxId:223] [49552] (60 PDB entries) 
Domain d1nifa2: 1nif A:167-340 [23049]
complexed with cu
Details for d1nifa2
PDB Entry: 1nif (more details), 1.7 Å
PDB Description: the structure of cu-nitrite reductase from achromobacter cycloclastes at five ph values, with nitrite bound and with type ii cu depleted
PDB Compounds: (A:) nitrite reductaseSCOPe Domain Sequences for d1nifa2:
Sequence; same for both SEQRES and ATOM records: (download)
>d1nifa2 b.6.1.3 (A:167-340) Nitrite reductase, NIR {Achromobacter cycloclastes [TaxId: 223]}
gqpltydkiyyvgeqdfyvpkdeagnykkyetpgeayedavkamrtltpthivfngavga
ltgdhaltaavgervlvvhsqanrdtrphligghgdyvwatgkfrnppdldqetwlipgg
tagaafytfrqpgvyayvnhnlieafelgaaghfkvtgewnddlmtsvvkpasm
SCOPe Domain Coordinates for d1nifa2:
Click to download the PDB-style file with coordinates for d1nifa2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1nifa2:
- d1nifa2 first appeared (with stable ids) in SCOP 1.55, called d1nif_2
- d1nifa2 appears in SCOPe 2.06
- d1nifa2 appears in the current release, SCOPe 2.08
 View in 3D
View in 3D