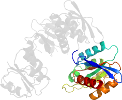
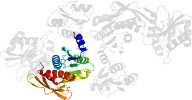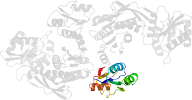Lineage for d1gqqb3 (1gqq B:107-321)
- Root: SCOPe 2.06

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.72: Ribokinase-like [53612] (3 superfamilies)
core: 3 layers: a/b/a; mixed beta-sheet of 8 strands, order 21345678, strand 7 is antiparallel to the rest
potential superfamily: members of this fold have similar functions but different ATP-binding sites
Superfamily c.72.2: MurD-like peptide ligases, catalytic domain [53623] (3 families) 
has extra strand located between strands 1 and 2
Family c.72.2.1: MurCDEF [53624] (5 proteins)
automatically mapped to Pfam PF08245
Protein UDP-N-acetylmuramate-alanine ligase MurC [82520] (2 species) 
Species Haemophilus influenzae [TaxId:727] [89776] (4 PDB entries) 
Domain d1gqqb3: 1gqq B:107-321 [83304]
Other proteins in same PDB: d1gqqa1, d1gqqa2, d1gqqb1, d1gqqb2
Details for d1gqqb3
PDB Entry: 1gqq (more details), 3.1 Å
PDB Description: murc - crystal structure of the apo-enzyme from haemophilus influenzae
PDB Compounds: (B:) udp-n-acetylmuramate-l-alanine ligaseSCOPe Domain Sequences for d1gqqb3:
Sequence, based on SEQRES records: (download)
>d1gqqb3 c.72.2.1 (B:107-321) UDP-N-acetylmuramate-alanine ligase MurC {Haemophilus influenzae [TaxId: 727]}
raqmlaeimrfrhgiavagthgkttttamismiytqakldptfvngglvksagknahlga
sryliaeadesdasflhlqpmvsvvtnmepdhmdtyegdfekmkatyvkflhnlpfygla
vmcaddpvlmelvpkvgrqvitygfseqadyriedyeqtgfqghytvicpnnerinvlln
vpgkhnalnataalavakeegianeailealadfq
Sequence, based on observed residues (ATOM records): (download)
>d1gqqb3 c.72.2.1 (B:107-321) UDP-N-acetylmuramate-alanine ligase MurC {Haemophilus influenzae [TaxId: 727]}
raqmlaeimrfrhgiavagthgkttttamismiytqakldptfvryliaeadeflhlqpm
vsvvtnmepfekmkatyvkflhnlpfyglavmcaddpvlmelvpkvgrqvitygfseqad
yriedyeqtgfqghytvicpnnerinvllnvpgkhnalnataalavakeegianeailea
ladfq
SCOPe Domain Coordinates for d1gqqb3:
Click to download the PDB-style file with coordinates for d1gqqb3.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1gqqb3:
- d1gqqb3 first appeared in SCOP 1.65
- d1gqqb3 appears in SCOPe 2.05
- d1gqqb3 appears in SCOPe 2.07
- d1gqqb3 appears in the current release, SCOPe 2.08

 View in 3D
View in 3D