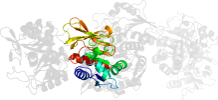
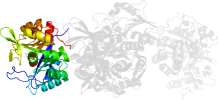
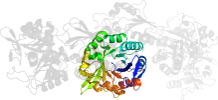Lineage for d1gpwd_ (1gpw D:)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.23: Flavodoxin-like [52171] (15 superfamilies)
3 layers, a/b/a; parallel beta-sheet of 5 strand, order 21345
Superfamily c.23.16: Class I glutamine amidotransferase-like [52317] (10 families) 
conserved positions of the oxyanion hole and catalytic nucleophile; different constituent families contain different additional structures
Family c.23.16.1: Class I glutamine amidotransferases (GAT) [52318] (12 proteins)
contains a catalytic Cys-His-Glu triad
Protein GAT subunit, HisH, (or domain) of imidazoleglycerolphosphate synthase HisF [69447] (4 species) 
Species Thermotoga maritima [TaxId:2336] [69448] (9 PDB entries) 
Domain d1gpwd_: 1gpw D: [65463]
Other proteins in same PDB: d1gpwa_, d1gpwc_, d1gpwe_
complexed with po4
Details for d1gpwd_
PDB Entry: 1gpw (more details), 2.4 Å
PDB Description: structural evidence for ammonia tunneling across the (beta/alpha)8 barrel of the imidazole glycerol phosphate synthase bienzyme complex.
PDB Compounds: (D:) amidotransferase hishSCOPe Domain Sequences for d1gpwd_:
Sequence; same for both SEQRES and ATOM records: (download)
>d1gpwd_ c.23.16.1 (D:) GAT subunit, HisH, (or domain) of imidazoleglycerolphosphate synthase HisF {Thermotoga maritima [TaxId: 2336]}
mrigiisvgpgnimnlyrgvkrasenfedvsielvesprndlydllfipgvghfgegmrr
lrendlidfvrkhvederyvvgvclgmqllfeeseeapgvkglsliegnvvklrsrrlph
mgwnevifkdtfpngyyyfvhtyravceeehvlgtteydgeifpsavrkgrilgfqfhpe
ksskigrkllekviecslsrr
SCOPe Domain Coordinates for d1gpwd_:
Click to download the PDB-style file with coordinates for d1gpwd_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1gpwd_:
- d1gpwd_ first appeared in SCOP 1.59
- d1gpwd_ appears in SCOPe 2.07

 View in 3D
View in 3D