SCOP 1.63 help
Overview
Structural Classification of Proteins — extended (SCOPe) is a database of protein structural relationships that extends the SCOP database. SCOP is a (mostly) manually curated ordering of domains from the majority of proteins of known structure in a hierarchy according to structural and evolutionary relationships. Development of SCOP (version 1) concluded with SCOP 1.75, released in June 2009. SCOPe extends the SCOP database, using a combination of manual curation and rigorously validated automated methods to classify many newer PDB structures. Our goal in implementing automation is to extend SCOPe to include structures that can be classified in the SCOP hierarchy, without sacrificing the reliability of the database that was developed through years of expert curation.
SCOPe also incorporates and updates the Astral compendium. Astral provides several databases and tools to aid in the analysis of the protein structures classified in SCOP, particularly through the use of their sequences. Here is a link to more info about Astral. New releases of Astral are derived from SCOPe.
The SCOPe website provides integrated access to data found in all releases of the SCOP and Astral databases that feature stable identifiers.
The SCOP hierarchy
- Applies to: SCOP version 1.55 through current release
- Reference: 5
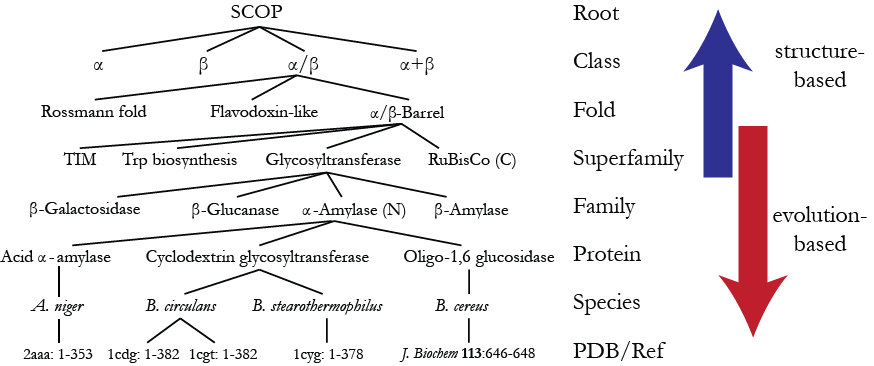
By analogy with taxonomy, SCOP was created as a hierarchy of several levels where the fundamental unit of classification is a domain in the experimentally determined protein structure. Starting at the bottom, the hierarchy of SCOP domains comprises the following levels:
- Species representing a distinct protein sequence and its naturally occurring or artificially created variants.
- Protein grouping together similar sequences of essentially the same functions that either originate from different biological species or represent different isoforms within the same species
- Family containing proteins with similar sequences but typically distinct functions
- Superfamily bridging together protein families with common functional and structural features inferred to be from a common evolutionary ancestor.
- Folds grouping structurally similar superfamilies.
- Classes based mainly on secondary structure content and organization.
Only the first seven classes are true classes. Because traditionally the aim of SCOP has been to classify every residue in the PDB, the remaining classes were maintained as placeholders to keep track of portions of a PDB structure that were not appropriate to include in SCOP. The SCOPe automated classification methods do not currently add entries to classes other than the first seven.
The multi-domain class is a special class. It contains chains with multiple domains that haven't been observed in a different context.
Changes to SCOP(e) design and size
- Applies to: SCOP version 1.55 through current release
- References: 1-5,8
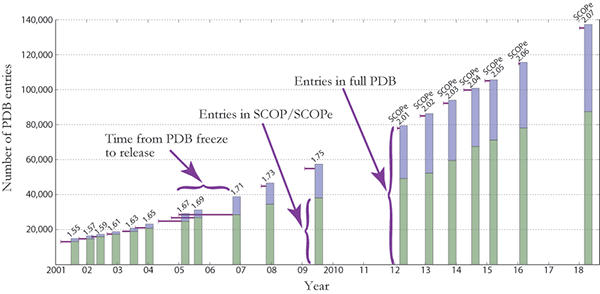
The figure below shows the number of structures in the PDB and
SCOP(e) at the time of
each SCOP(e) release. Extended
horizontal lines start at the freeze date for
each SCOP(e) release and show the
number of PDB entries available on that date. (The "freeze date"
is the last date for PDB entries to be released and still classified
in a given SCOP(e) release. Prior to
SCOP 1.73, all protein structures available on
the freeze date were manually classified.)
The figure below, adapted from
[3],
illustrates how SCOP(e)
has changed over the years. The length of the vertical line for
each release is proportional to the number of entries classified. The angle
of the blue baseline between releases reflects the degree of
divergence from comprehensive and fully manually curated
releases. The angle is based on the percentage of PDB entries on the
freeze date that were not classified in the release, and the extent
to which entries were manually classified.
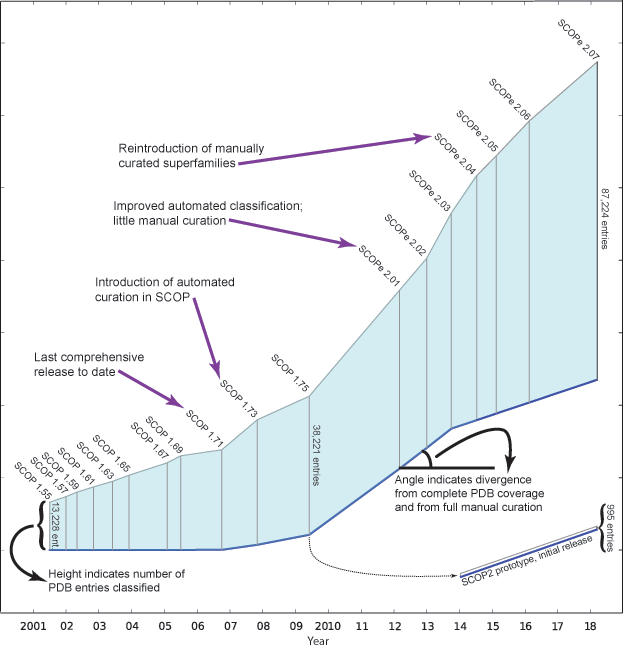
Note that since releases beyond SCOP 1.71 are not comprehensive, not all structurally characterized protein families and folds from the PDB are classified in these releases. Therefore, we caution against using later releases to (for example) analyze the rate at which new folds are being discovered.
Stable identifiers
- Applies to: SCOP version 1.55 through current release
- Reference: 8
The SCOPe database continues to support the same style of stable identifiers in use since SCOP 1.55. Identifiers are provided as an unambiguous way to link to each a SCOP or SCOPe entry and are stable across releases. See the sections on citing and linking to SCOPe and SCOP using stable identifiers.
-
sccs. SCOP(e) concise classification string. This is a dot notation used to concisely describe a SCOP(e) class, fold, superfamily, and family. For example, a.39.1.1 references the "Calbindin D9K" family, where "a" represents the class, "39" represents the fold, "1" represents the superfamily, and the last "1" represents the family.
-
sunid. SCOP(e) unique identifier. This is simply a number that may be used to reference any entry in the SCOP(e) hierarchy, from root to leaves (Fold, Superfamily, Family, etc.).
-
sid. Stable domain identifier. A 7-character sid consists of "d" followed by the 4-character PDB ID of the file of origin, the PDB chain ID ('_' if none, '.' if multiple as is the case in genetic domains), and a single character (usually an integer) if needed to specify the domain uniquely ('_' if not). Sids are currently all lower case, even when the chain letter is upper case. Example sids include d4akea1, d9hvpa_, and d1cph.1.
Both sunids and sccs identifiers are expected to remain stable across releases, except in cases where the classification changes substantially. For example, when nodes in the hierarchy (e.g. Superfamilies) are merged or split due to new evidence of evolutionary relationships, corresponding identifiers become obsolete and new sunids are introduced. If a domain is split, or the boundaries change substantially, new sid(s) and sunid(s) are assigned.
A history of changes between all consecutive SCOP(e) releases is available under the Stats & History tab, and a history of changes to each individual entry is shown at the bottom of the page describing that entry.
Astral documentation
- Applies to: SCOP version 1.55 through current release
- References: 11-13
The primary sources of Astral documention are the three references cited above and listed on the Help > References page. The Astral compendium is a collection of software and databases, partially derived from SCOP(e), that aid research into protein structure and evolution. Astral provides sequences and coordinate files for all SCOP(e) domains, as well as sequences for all PDB chains that are classified in SCOP(e). Chemically modified amino acids are translated back to the original sequence, and sequences are curated to eliminate errors resulting from the automated parsing of PDB files. Because the majority of sequences in the PDB are very similar to others, Astral provides representative subsets of proteins that span the set of classified protein structures or domains while alleviating bias toward well-studied proteins. The highest quality representative in each subset is chosen using AEROSPACI scores, which provide a numeric estimate of the quality and precision of crystallographically determined structures.
The figure below, taken from the
2004 NAR paper, gives a brief overview
of how Astral is created. New releases
of Astral are derived from SCOPe domain definitions.

Data flow in Astral: Primary data sources are shown in green. Primary Astral databases are shown in light yellow. Less commonly used resources are shown in darker yellow. Resources added more recently are outlined in light blue/grey. Using the RAF maps, four complete sequence sets are created for every domain in the first seven classes of the SCOP(e) database. Two sets (the genetic domain sets) include the genetic domain sequences described in the 2002 NAR paper, and the other two (the original-style sequence sets) use the prior method of splitting each multi-chain domain into multiple sequences. For each of these methodologies, one complete sequence set is derived from sequences in the PDB ATOM records, and another from sequences in the SEQRES records. The SEQRES sets (for both genetic domain and original-style methods) are used to derive representative subsets. Each set is fully compared against itself using BLAST, and subsets are created using three similarity criteria and various thresholds. Representatives are chosen according to AEROSPACI scores. PDB chain sequence sets are derived from the SEQRES records of every PDB chain in SCOP(e); selected subsets are created at 90-100% ID thresholds. PDB-style files are derived from the RAF maps and SCOP(e) domain definitions. At each new release of Astral, all non-redundant sequences from each SCOP(e) family and superfamily are aligned using MAFFT. A hidden Markov model (HMM) is created from the multiple sequence alignment for each family and superfamily using HMMER. These HMMs, and BLAST, are used to predict domains in the sequences of newly released PDB entries on a weekly basis. HMMs from the Pfam-A database are also used to predict domains in regions of the sequences not identified by HMMs derived from Astral. Unassigned regions of at least 20 consecutive residues are also predicted to be potential domains.
Documentation describing more recent updates (since the 2004 paper) can be found in the release notes, which are linked from the Downloads > Astral Sequences & Subsets page.
Searching SCOP and SCOPe
- Applies to: SCOP version 1.55 through current release
- Reference: 8
The website search bar at the top of each page supports keyword search, as well as searches by a number of different SCOP(e) identifiers.
View the Search examples page for further details on the syntax, as well as many examples.
Domain visualization
- Applies to: SCOP version 1.55 through current release
- Reference: 1
Thumbnails were generated for each domain using PyMOL, based on a viewing angle for each protein calculated using OVOP (Sverud O, MacCallum RM. 2003. Towards optimal views of proteins. Bioinformatics 19(7): 882-888 [PubMed]).
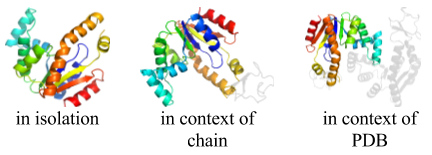
On the SCOPe website page showing information about each domain, thumbnails are displayed showing the domain in isolation, in the context of its chain, and in the context of its PDB structure. Links to other domains in the same chain, and in the same PDB structure, are below the corresponding thumbnail. To preview thumbnails from these other domains and get tool tip text with a short description, mouse over the links to the other domains. The figure to the right shows the thumbnails generated for the domain d4akea1.
Domain pages also include a JavaScript-based viewer that allows users to view and rotate domains in 3D without installing additional software. The 3D visualization domain visualization tool was built using JSmol, a Javascript-based viewer created by the Jmol project.
How to link to SCOP and SCOPe nodes
- Applies to: SCOP version 1.55 through current release
- References: 1,8
Because the SCOPe website is based on different technology from the older SCOP websites, old search paths (e.g., those using the search.cgi engine) are deprecated. URLs in the older style will be redirected approprately, so all old links to the Berkeley SCOP mirror should still work, including links to retrieve data that used to be on the Astral website (e.g., getseqs.cgi, spaci.cgi). We also support old-style links to all versions of SCOP with stable identifiers; for example, http://scop.berkeley.edu/scop/ links to SCOP 1.75, the last release of SCOP (version 1), while http://scop.berkeley.edu/scop-1.67/ links to SCOP version 1.67.
When making new links to SCOP or SCOPe data on this website, please use the new link formats below:
-
To link to a node the current release, use any
stable identifier (sunid,
sid or sccs).
The link formats are:
http://scop.berkeley.edu/sunid=xxxxx http://scop.berkeley.edu/sid=xxxxx http://scop.berkeley.edu/sccs=xxxxx
Linking by sunid is preferred because it is faster (sunid is a number, e.g. 46456; sids and sccs are both alphanumeric). Note that if the identifier you are linking to has become obsolete in the current release, the last release in which the identifier appears is shown, along with an explanation. -
To link to a node in a particular SCOP
or SCOPe
release, use the version number and sunid. The link format is:
http://scop.berkeley.edu/sunid=xxxxx&ver=yyyy
(versions are alphanumeric; e.g., 1.55 or 2.01).
You can also use this style of linking to link to nodes by sid or by sccs. -
To link to the root of a particular SCOP
or SCOPe
release, use the version number. The link format is:
http://scop.berkeley.edu/ver=yyyy
(versions are alphanumeric; e.g., 1.55 or 2.01). -
To link to information we have on a PDB
file, use its PDB code. The link
format is:
http://scop.berkeley.edu/pdb/code=xxxx
We plan for this link format to remain stable for all future versions of SCOPe unless there is a major change to the classification, even if we make changes to our underlying website technology.
How to cite SCOP and SCOPe entries
- Applies to: SCOP version 1.55 through current release
- References: 1,8
In the interest of scientific reproducibility, when you publish a manuscript or give a talk about data you obtained from SCOP or SCOPe, it's crucial to unambiguously refer to the data you used. Using these citation conventions will provide all the information needed for other researchers to accurately download the same data that you are working with. For related help, see the section on stable identifiers.
When citing a particular SCOPe or SCOP entry, we recommend the following conventions, depending on the level of the entry cited:
- Class: Use the complete SCOP(e)
version number, full name of the class, and
sunid. Examples:
- SCOPe 2.03-stable Class a: All alpha proteins [46456]
- SCOP 1.67 Class c: Alpha and beta proteins (a/b) [51349]
- Fold, Superfamily, or Family: Use the complete SCOP(e)
version number, sccs,
full name, and sunid. Examples:
- SCOPe 2.03-2013-12-12 Fold b.1: Immunoglobulin-like beta-sandwich [48725]
- SCOPe 2.03-stable Family c.2.1.1: Alcohol dehydrogenase-like, C-terminal domain [51736]
-
Protein or Species: Use the complete SCOP(e)
version number, full
name, sunid, and sccs. Examples:
- SCOPe 2.03-stable Protein Adenylate kinase [52554] (sccs c.37.1.1)
- SCOP 1.75 Species: Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932] [56639] (sccs e.5.1.1)
-
Domain. Use the complete SCOP(e)
version number, sid, sunid, and sccs. Examples:
- SCOPe 2.03-stable d1p0ca2 [87639] (sccs c.2.1.1)
- SCOPe 2.03-2013-12-12 d4cadj2 [229759] (sccs b.1.1.0)
Publications using SCOP, Astral or SCOPe data should also cite the appropriate references given on the Help > References page rather than just referring to the website.
Parseable files
- Applies to: SCOP version 1.55 through current release
- Reference: 8
Parseable files have been provided for each SCOP(e) release, available from the Downloads > Parseable Files & Software page. Each of these files has a header, starting with the '#' character, which includes release, version, and copyright information.
The list below shows the four parseable formats for SCOP(e) data. For information on downloading and parsing Astral subsets and RAF sequence maps, see the Astral help.
-
dir.des.scop(e).txt. - Descriptions for each node in the SCOP(e) hierarchy.
46457 cf a.1 - Globin-like 113449 px a.1.1.1 d1ux8a_ 1ux8 A: ^1 ^2 ^3 ^4 ^5
Five tab-delimited columns:
- sunid
- level: cl - class; cf - fold; sf - superfamily; fa - family; dm - protein; sp - species; px - domain
- sccs
- sid or "-"
- description
-
dir.cla.scop(e).txt - Full classification for each domain.
d1ux8a_ 1ux8 A: a.1.1.1 113449 cl=46456,cf=46457,sf=46458,fa=46459,dm=46460,sp=116748,px=113449 2gtld1 2gtl D:8-147 a.1.1.2 135658 cl=46456,cf=46457,sf=46458,fa=46463,dm=116758,sp=116759,px=135658 d1cph.1 1cph B:,A: g.1.1.1 43831 cl=56992,cf=56993,sf=56994,fa=56995,dm=56996,sp=56997,px=43831 ^1 ^2 ^3 ^4 ^5 ^6
Six tab-delimited columns:
- sid
- PDB ID
- description
- sccs
- sunid
- sunids of ancestor nodes in comma-delimited list, in the format "level=sunid". For level codes, see description for dir.des.scop(e).txt.
-
dir.hie.scop(e).txt - Children and parents for each node
0 - 46456,48724,51349,53931,56572,56835,56992,57942,58117,58231,58788 46457 46456 46458,46548 ^1 ^2 ^3
Three tab-delimited columns:
- sunid
- parent's sunid or "-"
- comma-separated list of children's sunids or "-"
-
dir.com.scop(e).txt - Human and machine-annotated comments
100067 ! complexed with cyn, hem, xe 63437 ! SQ Q10784 164742 ! automated match to d1s56b_ ! complexed with cyn, hem, na; mutant ^1 ^2 ^3
Two or more "!"-delimited columns:
- sunid
- first comment
- additional comments (one per column)
Plans for future website development
We are continuing to upgrade the website. Planned features include more sophisticated search functionality (including searches by sequence and by structure), use of AJAX to avoid full page reloads, and a SCOPe tree browser for more efficient display of results. We also plan to allow downloads of additional derived data (e.g., sequence sets and HMMs) for different clades in the tree.
Further help
The primary sources of documentation are the papers listed on the Help > References page.Please report bugs and usability issues to us at scope@compbio.berkeley.edu.
