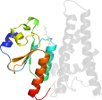Lineage for d1k0db1 (1k0d B:201-353)
- Root: SCOP 1.65

Class a: All alpha proteins [46456] (179 folds) 
Fold a.45: Glutathione S-transferase (GST), C-terminal domain [47615] (1 superfamily)
core: 4 helices; bundle, closed, left-handed twist; right-handed superhelix
Superfamily a.45.1: Glutathione S-transferase (GST), C-terminal domain [47616] (1 family) 
this domains follows the thioredoxin-like N-terminal domain
Family a.45.1.1: Glutathione S-transferase (GST), C-terminal domain [47617] (15 proteins) 
Protein Yeast prion protein ure2p, nitrogen regulation fragment [47641] (1 species)
similar to class phi enzymes
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [47642] (8 PDB entries) 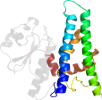
Domain d1k0db1: 1k0d B:201-353 [67932]
Other proteins in same PDB: d1k0da2, d1k0db2, d1k0dc2, d1k0dd2
Details for d1k0db1
PDB Entry: 1k0d (more details), 2.2 Å
PDB Description: Ure2p in Complex with Glutathione
SCOP Domain Sequences for d1k0db1:
Sequence, based on SEQRES records: (download)
>d1k0db1 a.45.1.1 (B:201-353) Yeast prion protein ure2p, nitrogen regulation fragment {Baker's yeast (Saccharomyces cerevisiae)}
lwsddladqsqinawlffqtsghapmigqalhfryfhsqkiasaverytdevrrvygvve
malaerrealvmeldtenaaaysagttpmsqsrffdypvwlvgdkltiadlafvpwnnvv
driginikiefpevykwtkhmmrrpavikalrg
Sequence, based on observed residues (ATOM records): (download)
>d1k0db1 a.45.1.1 (B:201-353) Yeast prion protein ure2p, nitrogen regulation fragment {Baker's yeast (Saccharomyces cerevisiae)}
lwsddladqsqinawlffqtsghapmigqalhfryfhsqkiasaverytdevrrvygvve
malaerrealvmdypvwlvgdkltiadlafvpwnnvvdriginikiefpevykwtkhmmr
rpavikalrg
SCOP Domain Coordinates for d1k0db1:
Click to download the PDB-style file with coordinates for d1k0db1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1k0db1:
- d1k0db1 first appeared in SCOP 1.59
- d1k0db1 appears in SCOP 1.63
- d1k0db1 appears in SCOP 1.67
- d1k0db1 appears in the current release, SCOPe 2.08

 View in 3D
View in 3D