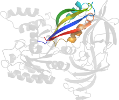
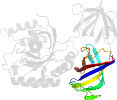Lineage for d1ijfa3 (1ijf A:2-240)
- Root: SCOP 1.67

Class c: Alpha and beta proteins (a/b) [51349] (130 folds) 
Fold c.37: P-loop containing nucleoside triphosphate hydrolases [52539] (1 superfamily)
3 layers: a/b/a, parallel or mixed beta-sheets of variable sizes
Superfamily c.37.1: P-loop containing nucleoside triphosphate hydrolases [52540] (22 families) 
division into families based on beta-sheet topologies
Family c.37.1.8: G proteins [52592] (37 proteins)
core: mixed beta-sheet of 6 strands, order 231456; strand 2 is antiparallel to the rest
Protein Elongation factor eEF-1alpha, N-terminal (G) domain [52631] (2 species)
eukaryotic and archaeal homologue of EF-Tu
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [52632] (4 PDB entries) 
Domain d1ijfa3: 1ijf A:2-240 [62491]
Other proteins in same PDB: d1ijfa1, d1ijfa2, d1ijfb_
complexed with gdp, mg
Details for d1ijfa3
PDB Entry: 1ijf (more details), 3 Å
PDB Description: Nucleotide exchange mechanisms in the eEF1A-eEF1Ba complex
SCOP Domain Sequences for d1ijfa3:
Sequence; same for both SEQRES and ATOM records: (download)
>d1ijfa3 c.37.1.8 (A:2-240) Elongation factor eEF-1alpha, N-terminal (G) domain {Baker's yeast (Saccharomyces cerevisiae)}
gkekshinvvvighvdsgkstttghliykcggidkrtiekfekeaaelgkgsfkyawvld
klkaerergitidialwkfetpkyqvtvidapghrdfiknmitgtsqadcailiiaggvg
efeagiskdgqtrehallaftlgvrqlivavnkmdsvkwdesrfqeivketsnfikkvgy
npktvpfvpisgwngdnmieattnapwykgweketkagvvkgktlleaidaieqpsrpt
SCOP Domain Coordinates for d1ijfa3:
Click to download the PDB-style file with coordinates for d1ijfa3.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1ijfa3:
- d1ijfa3 first appeared in SCOP 1.57
- d1ijfa3 appears in SCOP 1.65
- d1ijfa3 appears in SCOP 1.69
- d1ijfa3 appears in the current release, SCOPe 2.08
 View in 3D
View in 3D