Lineage for d1rypu_ (1ryp U:)
- Root: SCOP 1.55

Class d: Alpha and beta proteins (a+b) [53931] (184 folds) 
Fold d.153: N-terminal nucleophile aminohydrolases (Ntn hydrolases) [56234] (1 superfamily) 
Superfamily d.153.1: N-terminal nucleophile aminohydrolases (Ntn hydrolases) [56235] (5 families) 

Family d.153.1.4: Proteasome subunits [56251] (3 proteins) 
Protein Proteasome alpha subunit (non-catalytic) [56255] (2 species) 
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [56257] (3 PDB entries) 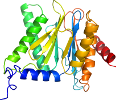
Domain d1rypu_: 1ryp U: [41945]
Other proteins in same PDB: d1ryp1_, d1ryp2_, d1ryph_, d1rypi_, d1rypj_, d1rypk_, d1rypl_, d1rypm_, d1rypn_, d1rypv_, d1rypw_, d1rypx_, d1rypy_, d1rypz_
Details for d1rypu_
PDB Entry: 1ryp (more details), 1.9 Å
PDB Description: crystal structure of the 20s proteasome from yeast at 2.4 angstroms resolution
SCOP Domain Sequences for d1rypu_:
Sequence; same for both SEQRES and ATOM records: (download)
>d1rypu_ d.153.1.4 (U:) Proteasome alpha subunit (non-catalytic) {Baker's yeast (Saccharomyces cerevisiae)}
gtgydlsnsvfspdgrnfqveyavkavengttsigikcndgvvfaveklitskllvpqkn
vkiqvvdrhigcvysglipdgrhlvnrgreeaasfkklyktpipipafadrlgqyvqaht
lynsvrpfgvstifggvdkngahlymlepsgsywgykgaatgkgrqsakaeleklvdhhp
eglsareavkqaakiiylahednkekdfeleiswcslsetnglhkfvkgdllqeaidfaq
kein
SCOP Domain Coordinates for d1rypu_:
Click to download the PDB-style file with coordinates for d1rypu_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1rypu_:
- d1rypu_ appears in SCOP 1.57
- d1rypu_ appears in the current release, SCOPe 2.08
 View in 3D View in 3DDomains from other chains: (mouse over for more information) d1ryp1_, d1ryp2_, d1rypa_, d1rypb_, d1rypc_, d1rypd_, d1rype_, d1rypf_, d1rypg_, d1ryph_, d1rypi_, d1rypj_, d1rypk_, d1rypl_, d1rypm_, d1rypn_, d1rypo_, d1rypp_, d1rypq_, d1rypr_, d1ryps_, d1rypt_, d1rypv_, d1rypw_, d1rypx_, d1rypy_, d1rypz_ |