Lineage for d1g6wc2 (1g6w C:100-200)
- Root: SCOPe 2.04

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.47: Thioredoxin fold [52832] (2 superfamilies)
core: 3 layers, a/b/a; mixed beta-sheet of 4 strands, order 4312; strand 3 is antiparallel to the rest
Superfamily c.47.1: Thioredoxin-like [52833] (24 families) 

Family c.47.1.5: Glutathione S-transferase (GST), N-terminal domain [52862] (19 proteins) 
Protein Yeast prion protein ure2p, nitrogen regulation fragment [52886] (1 species)
similar to class phi enzymes
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [52887] (8 PDB entries) 
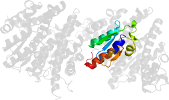
Domain d1g6wc2: 1g6w C:100-200 [33047]
Other proteins in same PDB: d1g6wa1, d1g6wb1, d1g6wc1, d1g6wd1
Details for d1g6wc2
PDB Entry: 1g6w (more details), 2.5 Å
PDB Description: crystal structure of the globular region of the prion protein ure2 from the yeast saccaromyces cerevisiae
PDB Compounds: (C:) ure2 proteinSCOPe Domain Sequences for d1g6wc2:
Sequence; same for both SEQRES and ATOM records: (download)
>d1g6wc2 c.47.1.5 (C:100-200) Yeast prion protein ure2p, nitrogen regulation fragment {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
sritkffqeqplegytlfshrsapngfkvaivlselgfhyntifldfnlgehrapefvsv
npnarvpalidhgmdnlsiwesgaillhlvnkyyketgnpl
SCOPe Domain Coordinates for d1g6wc2:
Click to download the PDB-style file with coordinates for d1g6wc2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1g6wc2:
- d1g6wc2 first appeared (with stable ids) in SCOP 1.55
- d1g6wc2 appears in SCOPe 2.03
- d1g6wc2 appears in SCOPe 2.05
- d1g6wc2 appears in the current release, SCOPe 2.08

 View in 3D
View in 3D