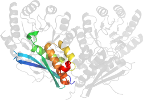
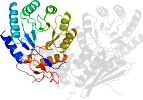
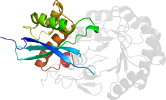Lineage for d3u9ib2 (3u9i B:131-371)
- Root: SCOPe 2.06

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.1: TIM beta/alpha-barrel [51350] (33 superfamilies)
contains parallel beta-sheet barrel, closed; n=8, S=8; strand order 12345678
the first seven superfamilies have similar phosphate-binding sites
Superfamily c.1.11: Enolase C-terminal domain-like [51604] (3 families) 
binds metal ion (magnesium or manganese) in conserved site inside barrel
N-terminal alpha+beta domain is common to this superfamily
Family c.1.11.0: automated matches [227196] (1 protein)
not a true family
Protein automated matches [226923] (74 species)
not a true protein
Species Roseiflexus sp. [TaxId:357808] [226247] (1 PDB entry) 
Domain d3u9ib2: 3u9i B:131-371 [217257]
Other proteins in same PDB: d3u9ia1, d3u9ib1
automated match to d1jpma1
complexed with so4
Details for d3u9ib2
PDB Entry: 3u9i (more details), 2.9 Å
PDB Description: The crystal structure of Mandelate racemase/muconate lactonizing enzyme from Roseiflexus sp.
PDB Compounds: (B:) Mandelate racemase/muconate lactonizing enzyme, C-terminal domain proteinSCOPe Domain Sequences for d3u9ib2:
Sequence, based on SEQRES records: (download)
>d3u9ib2 c.1.11.0 (B:131-371) automated matches {Roseiflexus sp. [TaxId: 357808]}
aatsletdvtittgsvtaaaraaqaivargvttikikigagdpdattirtmehdlariva
irdvaptarlildgncgytapdalrlldmlgvhgivpalfeqpvakddeeglrrltatrr
vpvaadesvasatdaarlarnaavdvlniklmkcgivealdiaaiartaglhlmiggmve
sllamtvsacfaagqggfrfvdldtplflaenpfdggmtyhggtidltlieaghgvtprs
p
Sequence, based on observed residues (ATOM records): (download)
>d3u9ib2 c.1.11.0 (B:131-371) automated matches {Roseiflexus sp. [TaxId: 357808]}
aatsletdvtittgsvtaaaraaqaivargvttikikigagttirtmehdlarivairdv
aptarlildgncgytapdalrlldmlgvhgivpalfeqpvakddeeglrrltatrrvpva
adesvasatdaarlarnaavdvlniklmkcgivealdiaaiartaglhlmiggmveslla
mtvsacfaagqggfrfvdldtplflaenpfdggmtyhggtidltlieaghgvtprsp
SCOPe Domain Coordinates for d3u9ib2:
Click to download the PDB-style file with coordinates for d3u9ib2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3u9ib2:
- d3u9ib2 first appeared in SCOPe 2.03
- d3u9ib2 appears in SCOPe 2.05
- d3u9ib2 appears in SCOPe 2.07
- d3u9ib2 appears in the current release, SCOPe 2.08
 View in 3D
View in 3D