Lineage for d4intv_ (4int V:)
- Root: SCOPe 2.06

Class d: Alpha and beta proteins (a+b) [53931] (385 folds) 
Fold d.153: Ntn hydrolase-like [56234] (2 superfamilies)
4 layers: alpha/beta/beta/alpha; has an unusual sheet-to-sheet packing
Superfamily d.153.1: N-terminal nucleophile aminohydrolases (Ntn hydrolases) [56235] (8 families) 
N-terminal residue provides two catalytic groups, nucleophile and proton donor
Family d.153.1.4: Proteasome subunits [56251] (4 proteins) 
Protein automated matches [190144] (11 species)
not a true protein
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [187078] (41 PDB entries) 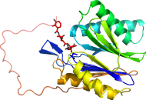
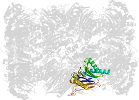
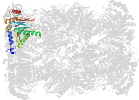
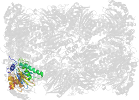
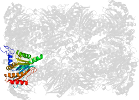
Domain d4intv_: 4int V: [202659]
Other proteins in same PDB: d4inta_, d4inte_, d4intf_, d4inti_, d4intj_, d4intk_, d4intl_, d4intm_, d4intn_, d4into_, d4ints_, d4intt_, d4intw_, d4intx_, d4inty_, d4intz_
automated match to d2zcyh_
complexed with 1g5
Details for d4intv_
PDB Entry: 4int (more details), 2.9 Å
PDB Description: Yeast 20S proteasome in complex with the vinyl sulfone LU122
PDB Compounds: (V:) Proteasome component PUP1SCOPe Domain Sequences for d4intv_:
Sequence; same for both SEQRES and ATOM records: (download)
>d4intv_ d.153.1.4 (V:) automated matches {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
ttivgvkfnngvviaadtrstqgpivadkncaklhrispkiwcagagtaadteavtqlig
snielhslytsreprvvsalqmlkqhlfkyqghigaylivagvdptgshlfsihahgstd
vgyylslgsgslaamavleshwkqdltkeeaiklasdaiqagiwndlgsgsnvdvcvmei
gkdaeylrnyltpnvreekqksykfprgttavlkesivnicd
SCOPe Domain Coordinates for d4intv_:
Click to download the PDB-style file with coordinates for d4intv_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d4intv_:
- d4intv_ first appeared in SCOPe 2.03
- d4intv_ appears in SCOPe 2.05
- d4intv_ appears in SCOPe 2.07
- d4intv_ appears in the current release, SCOPe 2.08
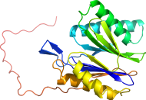
 View in 3D
View in 3D