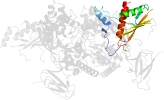
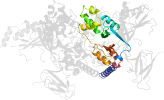
Lineage for d2qlve1 (2qlv E:161-247)
- Root: SCOPe 2.05

Class b: All beta proteins [48724] (176 folds) 
Fold b.1: Immunoglobulin-like beta-sandwich [48725] (31 superfamilies)
sandwich; 7 strands in 2 sheets; greek-key
some members of the fold have additional strands
Superfamily b.1.18: E set domains [81296] (24 families) 
"Early" Ig-like fold families possibly related to the immunoglobulin and/or fibronectin type III superfamilies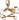
Family b.1.18.21: AMPK-beta glycogen binding domain-like [158886] (4 proteins)
lacks the N-terminal strand (A) and contains a beta-hairpin insertion in the C-terminal strand (G)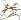
Protein SIP2 [158889] (1 species) 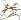
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [158890] (1 PDB entry)
Uniprot P34164 161-247
Domain d2qlve1: 2qlv E:161-247 [150870]
Other proteins in same PDB: d2qlva1, d2qlvb2, d2qlvc1, d2qlvc2, d2qlvd_, d2qlve2, d2qlvf1, d2qlvf2
automated match to d2qlvb1
Details for d2qlve1
PDB Entry: 2qlv (more details), 2.6 Å
PDB Description: crystal structure of the heterotrimer core of the s. cerevisiae ampk homolog snf1
PDB Compounds: (E:) Protein SIP2SCOPe Domain Sequences for d2qlve1:
Sequence; same for both SEQRES and ATOM records: (download)
>d2qlve1 b.1.18.21 (E:161-247) SIP2 {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
slmvpveirwqqggskvyvtgsftkwrkmiglipdsdnngsfhvklrllpgthrfrfivd
nelrvsdflptatdqmgnfvnyievrq
SCOPe Domain Coordinates for d2qlve1:
Click to download the PDB-style file with coordinates for d2qlve1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d2qlve1:
- d2qlve1 first appeared in SCOP 1.75
- d2qlve1 appears in SCOPe 2.04
- d2qlve1 appears in SCOPe 2.06
- d2qlve1 appears in the current release, SCOPe 2.08

 View in 3D
View in 3D