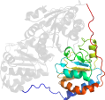
Lineage for Protein: Pyruvate decarboxylase
- Root: SCOP 1.67

Class c: Alpha and beta proteins (a/b) [51349] (130 folds) 
Fold c.31: DHS-like NAD/FAD-binding domain [52466] (1 superfamily)
3 layers: a/b/a; parallel beta-sheet of 6 strands, order 321456; Rossmann-like
Superfamily c.31.1: DHS-like NAD/FAD-binding domain [52467] (5 families) 
binds cofactor molecules in the opposite direction than classical Rossmann fold
Family c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain [52475] (7 proteins)
N-terminal domain is Pyr module, and C-terminal domain is PP module of thiamin diphosphate-binding fold
Protein Pyruvate decarboxylase [52478] (3 species)
rudiment domain with a variant fold, lacks FAD-binding
Species:

Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [52480] (2 PDB entries) - Domains for 1pvd:
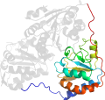
Domain d1pvda1: 1pvd A:182-360 [31737]
Other proteins in same PDB: d1pvda2, d1pvda3, d1pvdb2, d1pvdb3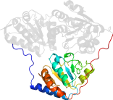
Domain d1pvdb1: 1pvd B:182-360 [31738]
Other proteins in same PDB: d1pvda2, d1pvda3, d1pvdb2, d1pvdb3
complexed with mg, tpp
- Domains for 1qpb:
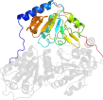
Domain d1qpba1: 1qpb A:182-360 [31739]
Other proteins in same PDB: d1qpba2, d1qpba3, d1qpbb2, d1qpbb3
Domain d1qpbb1: 1qpb B:182-360 [31740]
Other proteins in same PDB: d1qpba2, d1qpba3, d1qpbb2, d1qpbb3
- Domains for 1pvd:

Brewer's yeast (Saccharomyces), uvarum strain [TaxId:230603] [52479] (1 PDB entry) 
Zymomonas mobilis [TaxId:542] [52481] (1 PDB entry) - Domains for 1zpd:
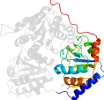
Domain d1zpda1: 1zpd A:188-362 [31741]
Other proteins in same PDB: d1zpda2, d1zpda3, d1zpdb2, d1zpdb3, d1zpde2, d1zpde3, d1zpdf2, d1zpdf3
Domain d1zpdb1: 1zpd B:188-362 [31742]
Other proteins in same PDB: d1zpda2, d1zpda3, d1zpdb2, d1zpdb3, d1zpde2, d1zpde3, d1zpdf2, d1zpdf3
Domain d1zpde1: 1zpd E:188-362 [31743]
Other proteins in same PDB: d1zpda2, d1zpda3, d1zpdb2, d1zpdb3, d1zpde2, d1zpde3, d1zpdf2, d1zpdf3
Domain d1zpdf1: 1zpd F:188-362 [31744]
Other proteins in same PDB: d1zpda2, d1zpda3, d1zpdb2, d1zpdb3, d1zpde2, d1zpde3, d1zpdf2, d1zpdf3
complexed with cit, dpx, mg
- Domains for 1zpd:
More info for Protein Pyruvate decarboxylase from c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain
Timeline for Protein Pyruvate decarboxylase from c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain:
- Protein Pyruvate decarboxylase from c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain first appeared (with stable ids) in SCOP 1.55
- Protein Pyruvate decarboxylase from c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain appears in SCOP 1.65
- Protein Pyruvate decarboxylase from c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain appears in SCOP 1.69
- Protein Pyruvate decarboxylase from c.31.1.3: Pyruvate oxidase and decarboxylase, middle domain appears in the current release, SCOPe 2.08