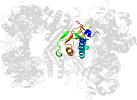
Lineage for d1gpmd2 (1gpm D:3-207)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.23: Flavodoxin-like [52171] (15 superfamilies)
3 layers, a/b/a; parallel beta-sheet of 5 strand, order 21345
Superfamily c.23.16: Class I glutamine amidotransferase-like [52317] (10 families) 
conserved positions of the oxyanion hole and catalytic nucleophile; different constituent families contain different additional structures
Family c.23.16.1: Class I glutamine amidotransferases (GAT) [52318] (12 proteins)
contains a catalytic Cys-His-Glu triad
Protein GMP synthetase [52319] (2 species) 
Species Escherichia coli [TaxId:562] [52320] (1 PDB entry) 
Domain d1gpmd2: 1gpm D:3-207 [31408]
Other proteins in same PDB: d1gpma1, d1gpma3, d1gpmb1, d1gpmb3, d1gpmc1, d1gpmc3, d1gpmd1, d1gpmd3
complexed with amp, cit, mg, po4, pop
Details for d1gpmd2
PDB Entry: 1gpm (more details), 2.2 Å
PDB Description: escherichia coli gmp synthetase complexed with amp and pyrophosphate
PDB Compounds: (D:) gmp synthetaseSCOPe Domain Sequences for d1gpmd2:
Sequence; same for both SEQRES and ATOM records: (download)
>d1gpmd2 c.23.16.1 (D:3-207) GMP synthetase {Escherichia coli [TaxId: 562]}
enihkhrilildfgsqytqlvarrvrelgvycelwawdvteaqirdfnpsgiilsggpes
tteensprapqyvfeagvpvfgvcygmqtmamqlgghveasnerefgyaqvevvndsalv
rgiedaltadgkplldvwmshgdkvtaipsdfitvastescpfaimaneekrfygvqfhp
evthtrqgmrmlerfvrdicqceal
SCOPe Domain Coordinates for d1gpmd2:
Click to download the PDB-style file with coordinates for d1gpmd2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1gpmd2:
- d1gpmd2 first appeared (with stable ids) in SCOP 1.55
- d1gpmd2 appears in SCOPe 2.07

 View in 3D
View in 3D