Lineage for d1jdca1 (1jdc A:358-418)
- Root: SCOPe 2.08

Class b: All beta proteins [48724] (180 folds) 
Fold b.71: Glycosyl hydrolase domain [51010] (1 superfamily)
folded sheet; greek-key
Superfamily b.71.1: Glycosyl hydrolase domain [51011] (6 families) 
this domain is C-terminal to the catalytic beta/alpha barrel domain
Family b.71.1.1: alpha-Amylases, C-terminal beta-sheet domain [51012] (22 proteins)
this domain follows the catalytic beta/alpha barrel domain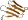
Protein G4-amylase (1,4-alpha-D-glucan maltotetrahydrolase) [51038] (1 species)
single beta-sheet; probable result of a decay of the common-fold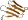
Species Pseudomonas stutzeri [TaxId:316] [51039] (9 PDB entries) 
Domain d1jdca1: 1jdc A:358-418 [27789]
Other proteins in same PDB: d1jdca2
complexed with ca; mutant
heterogeneous fold; applies to domains that adopt a different fold than the exemplar domain but has similar sequence and number of secondary structures
Details for d1jdca1
PDB Entry: 1jdc (more details), 1.9 Å
PDB Description: mutant (e219q) maltotetraose-forming exo-amylase cocrystallized with maltotetraose (crystal type 1)
PDB Compounds: (A:) 1,4-alpha maltotetrahydrolaseSCOPe Domain Sequences for d1jdca1:
Sequence; same for both SEQRES and ATOM records: (download)
>d1jdca1 b.71.1.1 (A:358-418) G4-amylase (1,4-alpha-D-glucan maltotetrahydrolase) {Pseudomonas stutzeri [TaxId: 316]}
radsaisfhsgysglvatvsgsqqtlvvalnsdlgnpgqvasgsfseavnasngqvrvwr
s
SCOPe Domain Coordinates for d1jdca1:
Click to download the PDB-style file with coordinates for d1jdca1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1jdca1:
- d1jdca1 first appeared (with stable ids) in SCOP 1.55, called d1jdc_1
- d1jdca1 appears in SCOPe 2.07

 View in 3D
View in 3D