Lineage for d1epme_ (1epm E:)
- Root: SCOPe 2.08

Class b: All beta proteins [48724] (180 folds) 
Fold b.50: Acid proteases [50629] (1 superfamily)
barrel, closed; n=6, S=10, complex topology
Superfamily b.50.1: Acid proteases [50630] (4 families) 
Family b.50.1.2: Pepsin-like [50646] (11 proteins)
duplication: consists of two similar barrel domains
N-terminal: barrel, partly opened; n*=6, S*=10
Protein Endothiapepsin [50647] (1 species) 
Species Chestnut blight fungus (Endothia parasitica) [TaxId:5116] [50648] (538 PDB entries) 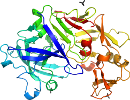
Domain d1epme_: 1epm E: [26781]
complexed with so4
Details for d1epme_
PDB Entry: 1epm (more details), 1.6 Å
PDB Description: a structural comparison of 21 inhibitor complexes of the aspartic proteinase from endothia parasitica
PDB Compounds: (E:) endothiapepsinSCOPe Domain Sequences for d1epme_:
Sequence; same for both SEQRES and ATOM records: (download)
>d1epme_ b.50.1.2 (E:) Endothiapepsin {Chestnut blight fungus (Endothia parasitica) [TaxId: 5116]}
stgsatttpidslddayitpvqigtpaqtlnldfdtgssdlwvfssettasevdgqtiyt
psksttakllsgatwsisygdgssssgdvytdtvsvggltvtgqavesakkvsssfteds
tidgllglafstlntvsptqqktffdnakasldspvftadlgyhapgtynfgfidttayt
gsitytavstkqgfwewtstgyavgsgtfkstsidgiadtgttllylpatvvsaywaqvs
gakssssvggyvfpcsatlpsftfgvgsarivipgdyidfgpistgssscfggiqssagi
ginifgdvalkaafvvfngattptlgfask
SCOPe Domain Coordinates for d1epme_:
Click to download the PDB-style file with coordinates for d1epme_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d1epme_:
- d1epme_ first appeared (with stable ids) in SCOP 1.55
- d1epme_ appears in SCOPe 2.07

 View in 3D
View in 3D