Lineage for d2zitc3 (2zit C:482-560)
- Root: SCOPe 2.04

Class d: Alpha and beta proteins (a+b) [53931] (380 folds) 
Fold d.58: Ferredoxin-like [54861] (59 superfamilies)
alpha+beta sandwich with antiparallel beta-sheet; (beta-alpha-beta)x2
Superfamily d.58.11: EF-G C-terminal domain-like [54980] (4 families) 
Family d.58.11.0: automated matches [254210] (1 protein)
not a true family
Protein automated matches [254469] (2 species)
not a true protein
Species Baker's yeast (Saccharomyces cerevisiae) [TaxId:4932] [255012] (4 PDB entries) 
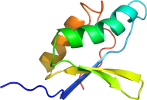
Domain d2zitc3: 2zit C:482-560 [239071]
Other proteins in same PDB: d2zita1, d2zita2, d2zita4, d2zitb_, d2zitc1, d2zitc2, d2zitc4, d2zitd_, d2zite1, d2zite2, d2zite4, d2zitf_
automated match to d1n0ua4
protein/RNA complex; complexed with nad
Details for d2zitc3
PDB Entry: 2zit (more details), 3 Å
PDB Description: structure of the eef2-exoa-nad+ complex
PDB Compounds: (C:) Elongation factor 2SCOPe Domain Sequences for d2zitc3:
Sequence; same for both SEQRES and ATOM records: (download)
>d2zitc3 d.58.11.0 (C:482-560) automated matches {Baker's yeast (Saccharomyces cerevisiae) [TaxId: 4932]}
kfsvspvvqvavevknandlpklveglkrlsksdpcvltymsesgehivagtgelhleic
lqdlehdhagvplkisppv
SCOPe Domain Coordinates for d2zitc3:
Click to download the PDB-style file with coordinates for d2zitc3.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d2zitc3:
- d2zitc3 is new in SCOPe 2.04-stable
- d2zitc3 appears in SCOPe 2.05
- d2zitc3 appears in the current release, SCOPe 2.08
 View in 3D View in 3DDomains from same chain: (mouse over for more information) d2zitc1, d2zitc2, d2zitc4, d2zitc5 |