Lineage for d3fmbb1 (3fmb B:1-99)
- Root: SCOPe 2.08

Class d: Alpha and beta proteins (a+b) [53931] (396 folds) 
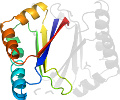
Fold d.58: Ferredoxin-like [54861] (62 superfamilies)
alpha+beta sandwich with antiparallel beta-sheet; (beta-alpha-beta)x2
Superfamily d.58.4: Dimeric alpha+beta barrel [54909] (24 families) 
dimerizes through the beta-sheet; forms beta-sheet barrel, closed (n=8, S=12); dimers may assemble in higher oligomers
Family d.58.4.0: automated matches [191316] (1 protein)
not a true family
Protein automated matches [190081] (33 species)
not a true protein
Species Bacteroides fragilis [TaxId:272559] [232176] (1 PDB entry) 
Domain d3fmbb1: 3fmb B:1-99 [232178]
Other proteins in same PDB: d3fmba2, d3fmbb2
automated match to d3bgua_
complexed with edo, so4
Details for d3fmbb1
PDB Entry: 3fmb (more details), 1.85 Å
PDB Description: crystal structure of dimeric protein of unknown function and ferredoxin-like fold (yp_212648.1) from bacteroides fragilis nctc 9343 at 1.85 a resolution
PDB Compounds: (B:) Dimeric protein of unknown function and ferredoxin-like foldSCOPe Domain Sequences for d3fmbb1:
Sequence; same for both SEQRES and ATOM records: (download)
>d3fmbb1 d.58.4.0 (B:1-99) automated matches {Bacteroides fragilis [TaxId: 272559]}
mvkhivlfklrddvpveeklvvmnsfkeaiealpakisvirkievglnmnpgetwnialy
sefdnlddvkfyathpehvaagkilaetkesracvdyef
SCOPe Domain Coordinates for d3fmbb1:
Click to download the PDB-style file with coordinates for d3fmbb1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3fmbb1:
- d3fmbb1 first appeared in SCOPe 2.03, called d3fmbb_
- d3fmbb1 appears in SCOPe 2.07

 View in 3D
View in 3D