Lineage for d4amvc2 (4amv C:1-240)
- Root: SCOPe 2.08

Class d: Alpha and beta proteins (a+b) [53931] (396 folds) 
Fold d.153: Ntn hydrolase-like [56234] (2 superfamilies)
4 layers: alpha/beta/beta/alpha; has an unusual sheet-to-sheet packing
Superfamily d.153.1: N-terminal nucleophile aminohydrolases (Ntn hydrolases) [56235] (8 families) 
N-terminal residue provides two catalytic groups, nucleophile and proton donor
Family d.153.1.1: Class II glutamine amidotransferases [56236] (6 proteins)
has slightly different topology than other families do
Protein Glucosamine 6-phosphate synthase, N-terminal domain [56237] (1 species) 
Species Escherichia coli [TaxId:562] [56238] (7 PDB entries)
Uniprot P17169 1-238
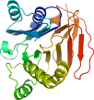
Domain d4amvc2: 4amv C:1-240 [219061]
Other proteins in same PDB: d4amva1, d4amvb1, d4amvc1
complexed with f6r
Details for d4amvc2
PDB Entry: 4amv (more details), 2.05 Å
PDB Description: e.coli glucosamine-6p synthase in complex with fructose-6p
PDB Compounds: (C:) glucosamine--fructose-6-phosphate aminotransferase [isomer izing]SCOPe Domain Sequences for d4amvc2:
Sequence; same for both SEQRES and ATOM records: (download)
>d4amvc2 d.153.1.1 (C:1-240) Glucosamine 6-phosphate synthase, N-terminal domain {Escherichia coli [TaxId: 562]}
cgivgaiaqrdvaeilleglrrleyrgydsaglavvdaeghmtrlrrlgkvqmlaqaaee
hplhggtgiahtrwathgepsevnahphvsehivvvhngiienheplreelkargytfvs
etdteviahlvnwelkqggtlreavlraipqlrgaygtvimdsrhpdtllaarsgsplvi
glgmgenfiasdqlallpvtrrfifleegdiaeitrrsvnifdktgaevkrqdiesnlqy
SCOPe Domain Coordinates for d4amvc2:
Click to download the PDB-style file with coordinates for d4amvc2.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d4amvc2:
- d4amvc2 first appeared in SCOPe 2.03
- d4amvc2 appears in SCOPe 2.07
 View in 3D View in 3DDomains from other chains: (mouse over for more information) d4amva1, d4amva2, d4amvb1, d4amvb2 |