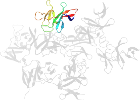Lineage for d3c0dg_ (3c0d G:)
- Root: SCOPe 2.08

Class b: All beta proteins [48724] (180 folds) 
Fold b.33: ISP domain [50021] (1 superfamily)
consists of two all-beta subdomains: conserved small domain has a rubredoxin-like fold; larger domain consists of 6 beta-stands packed in either sandwich of two 3-stranded sheets or closed barrel (n=6; S=8)
Superfamily b.33.1: ISP domain [50022] (4 families) 
Family b.33.1.3: NirD-like [158991] (1 protein)
retains the common fold but lacks the Fe-S cluster
automatically mapped to Pfam PF13806
Protein NADH-nitrite reductase small subunit NirD [158992] (3 species) 
Species Vibrio parahaemolyticus [TaxId:670] [158995] (1 PDB entry)
Uniprot Q87HB1 4-111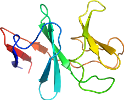
Domain d3c0dg_: 3c0d G: [155825]
Other proteins in same PDB: d3c0da2
automated match to d3c0da1
Details for d3c0dg_
PDB Entry: 3c0d (more details), 2.4 Å
PDB Description: crystal structure of the putative nitrite reductase nadph (small subunit) oxidoreductase protein q87hb1. northeast structural genomics consortium target vpr162
PDB Compounds: (G:) Putative nitrite reductase NADPH (Small subunit) oxidoreductase proteinSCOPe Domain Sequences for d3c0dg_:
Sequence; same for both SEQRES and ATOM records: (download)
>d3c0dg_ b.33.1.3 (G:) NADH-nitrite reductase small subunit NirD {Vibrio parahaemolyticus [TaxId: 670]}
ltkvklcqlddlmpfigatvliegervalfyipdsgvyavqdwdpigkayvmsrgivgdi
ngemcvasplykqhfslksgqcledeahclktwrvtvddnqvcyla
SCOPe Domain Coordinates for d3c0dg_:
Click to download the PDB-style file with coordinates for d3c0dg_.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d3c0dg_:
- d3c0dg_ first appeared in SCOP 1.75, called d3c0dg1
- d3c0dg_ appears in SCOPe 2.07
 View in 3D
View in 3D