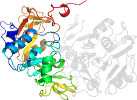Lineage for d2ahwa1 (2ahw A:284-529)
- Root: SCOPe 2.08

Class c: Alpha and beta proteins (a/b) [51349] (148 folds) 
Fold c.124: NagB/RpiA/CoA transferase-like [100949] (1 superfamily)
core: 3 layers: a/b/a; parallel or mixed beta-sheet of 6 strands, order 321456
Superfamily c.124.1: NagB/RpiA/CoA transferase-like [100950] (9 families) 

Family c.124.1.3: CoA transferase beta subunit-like [74657] (4 proteins)
catalytic subunit: similar active site structure to the NagB and RpiA families; mixed beta-sheet of 7 strands, order 4321567; strand 3 is antiparallel to the rest
Protein Putative enzyme YdiF C-terminal domain [142194] (1 species) 
Species Escherichia coli [TaxId:562] [142195] (3 PDB entries)
Uniprot P37766 283-529
Domain d2ahwa1: 2ahw A:284-529 [126801]
Other proteins in same PDB: d2ahwa2, d2ahwb2, d2ahwc2, d2ahwd2
automated match to d2ahua1
complexed with coa
Details for d2ahwa1
PDB Entry: 2ahw (more details), 2.15 Å
PDB Description: Crystal Structure of Acyl-CoA transferase from E. coli O157:H7 (YdiF)-thioester complex with CoA- 2
PDB Compounds: (A:) putative enzyme YdiFSCOPe Domain Sequences for d2ahwa1:
Sequence, based on SEQRES records: (download)
>d2ahwa1 c.124.1.3 (A:284-529) Putative enzyme YdiF C-terminal domain {Escherichia coli [TaxId: 562]}
plnqrklvarralfemrkgavgnvgvgiadgiglvareegcaddfiltvetgpiggitsq
giafganvntraildmtsqfdfyhgggldvcylsfaevdqhgnvgvhkfngkimgtggfi
disatskkiifcgtltagslkteiadgklnivqegrvkkfirelpeitfsgkialergld
vryiteravftlkedglhlieiapgvdlqkdildkmdftpvispelklmderlfidaamg
fvlpea
Sequence, based on observed residues (ATOM records): (download)
>d2ahwa1 c.124.1.3 (A:284-529) Putative enzyme YdiF C-terminal domain {Escherichia coli [TaxId: 562]}
plnqrklvarralfemrkgavgnvgvgiadgiglvareegcaddfiltvetgpiggitan
vntraildmtsqfdfyhgggldvcylsfaevdqhgnvgvhkfngkimgtggfidisatsk
kiifcgtltagslkteiadgklnivqegrvkkfirelpeitfsgkialergldvryiter
avftlkedglhlieiapgvdlqkdildkmdftpvispelklmderlfidaamgfvlpea
SCOPe Domain Coordinates for d2ahwa1:
Click to download the PDB-style file with coordinates for d2ahwa1.
(The format of our PDB-style files is described here.)
(The format of our PDB-style files is described here.)
Timeline for d2ahwa1:
- d2ahwa1 first appeared in SCOP 1.73
- d2ahwa1 appears in SCOPe 2.07

 View in 3D
View in 3D